You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000015_02842
You are here: Home > Sequence: MGYG000000015_02842
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterobacter mori | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Enterobacter; Enterobacter mori | |||||||||||
| CAZyme ID | MGYG000000015_02842 | |||||||||||
| CAZy Family | GH1 | |||||||||||
| CAZyme Description | 6-phospho-beta-glucosidase GmuD | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 303381; End: 304808 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 7 | 461 | 5.4e-119 | 0.9790209790209791 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2723 | BglB | 8.86e-152 | 9 | 460 | 5 | 449 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| pfam00232 | Glyco_hydro_1 | 1.13e-111 | 5 | 460 | 2 | 447 | Glycosyl hydrolase family 1. |
| PRK13511 | PRK13511 | 5.25e-87 | 7 | 466 | 4 | 468 | 6-phospho-beta-galactosidase; Provisional |
| PRK09589 | celA | 1.16e-64 | 57 | 460 | 62 | 468 | 6-phospho-beta-glucosidase; Reviewed |
| PRK15014 | PRK15014 | 1.56e-58 | 4 | 460 | 2 | 469 | 6-phospho-beta-glucosidase BglA; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QWC66176.1 | 0.0 | 1 | 475 | 1 | 475 |
| QXM21922.1 | 0.0 | 1 | 475 | 1 | 475 |
| BBT92058.1 | 0.0 | 1 | 475 | 1 | 475 |
| QCZ40031.1 | 0.0 | 1 | 475 | 1 | 475 |
| QPP01055.1 | 0.0 | 1 | 475 | 1 | 475 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4B3K_A | 1.02e-127 | 6 | 474 | 1 | 470 | Family1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3K_B Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3K_C Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3K_D Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3K_E Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3K_F Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_A Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_B Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_C Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_D Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_E Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes],4B3L_F Family 1 6-phospho-beta-D glycosidase from Streptococcus pyogenes [Streptococcus pyogenes] |
| 5FOO_A | 5.99e-127 | 6 | 474 | 2 | 471 | 6-phospho-beta-glucosidase[Streptococcus pyogenes M1 GAS],5FOO_B 6-phospho-beta-glucosidase [Streptococcus pyogenes M1 GAS],5FOO_C 6-phospho-beta-glucosidase [Streptococcus pyogenes M1 GAS],5FOO_D 6-phospho-beta-glucosidase [Streptococcus pyogenes M1 GAS],5FOO_E 6-phospho-beta-glucosidase [Streptococcus pyogenes M1 GAS],5FOO_F 6-phospho-beta-glucosidase [Streptococcus pyogenes M1 GAS] |
| 5NAV_A | 5.17e-119 | 4 | 460 | 16 | 471 | Crystalstructure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAV_B Crystal structure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAV_C Crystal structure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAV_D Crystal structure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAV_E Crystal structure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAV_F Crystal structure of the double mutant (Cys211Ser/Cys292Ser) 6-phospho-b-D-glucosidase from Lactobacillus plantarum [Lactiplantibacillus plantarum] |
| 5NAQ_A | 2.07e-118 | 4 | 460 | 16 | 471 | Crystalstructure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAQ_B Crystal structure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAQ_C Crystal structure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAQ_D Crystal structure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAQ_E Crystal structure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum],5NAQ_F Crystal structure of native 6-phospho-glucosidase LpBgl from Lactobacillus plantarum [Lactiplantibacillus plantarum] |
| 6Z1H_A | 5.35e-82 | 2 | 460 | 5 | 444 | ChainA, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1H_B Chain B, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1M_A Chain A, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_B Chain B, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_C Chain C, Ancestral reconstructed glycosidase [synthetic construct] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O05508 | 2.15e-131 | 3 | 460 | 4 | 459 | 6-phospho-beta-glucosidase GmuD OS=Bacillus subtilis (strain 168) OX=224308 GN=gmuD PE=1 SV=1 |
| P26208 | 6.26e-68 | 5 | 460 | 3 | 442 | Beta-glucosidase A OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=bglA PE=1 SV=1 |
| P10482 | 5.22e-63 | 6 | 459 | 3 | 448 | Beta-glucosidase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=bglA PE=3 SV=1 |
| Q97EZ2 | 8.22e-63 | 10 | 466 | 6 | 466 | 6-phospho-beta-galactosidase OS=Clostridium acetobutylicum (strain ATCC 824 / DSM 792 / JCM 1419 / LMG 5710 / VKM B-1787) OX=272562 GN=lacG PE=3 SV=1 |
| B3W7I3 | 9.09e-61 | 8 | 466 | 5 | 472 | 6-phospho-beta-galactosidase OS=Lacticaseibacillus casei (strain BL23) OX=543734 GN=lacG PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
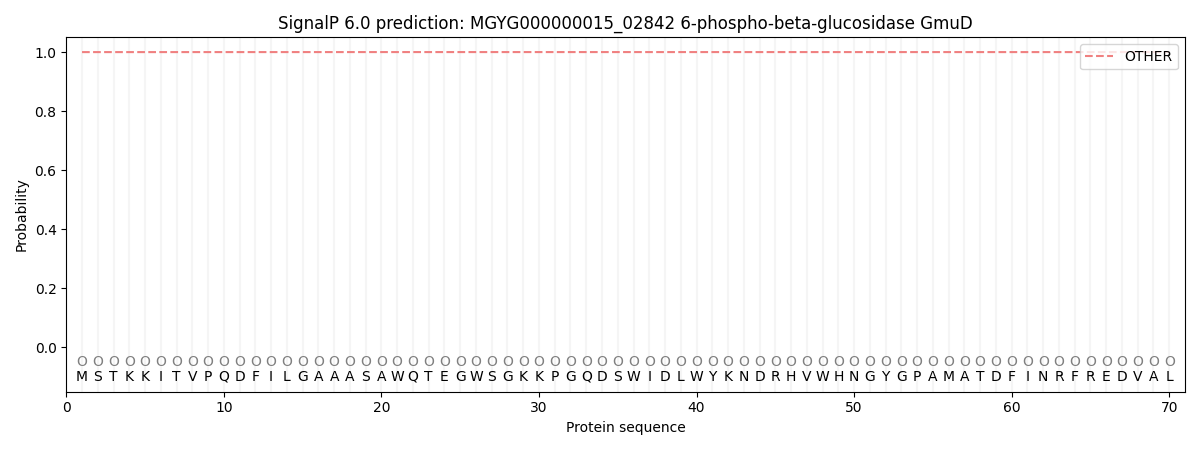
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000072 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
