You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000032_02833
You are here: Home > Sequence: MGYG000000032_02833
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Hungatella effluvii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Hungatella; Hungatella effluvii | |||||||||||
| CAZyme ID | MGYG000000032_02833 | |||||||||||
| CAZy Family | GH39 | |||||||||||
| CAZyme Description | Beta-xylosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 302911; End: 304428 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH39 | 16 | 439 | 2.7e-147 | 0.988399071925754 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01229 | Glyco_hydro_39 | 1.32e-129 | 22 | 482 | 20 | 487 | Glycosyl hydrolases family 39. |
| COG3664 | XynB | 1.84e-67 | 32 | 485 | 1 | 427 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNO18547.1 | 5.23e-234 | 4 | 499 | 10 | 507 |
| BCZ45334.1 | 1.48e-214 | 4 | 497 | 3 | 497 |
| VCV24167.1 | 1.26e-213 | 4 | 499 | 13 | 514 |
| CBL10149.1 | 2.54e-213 | 4 | 499 | 13 | 514 |
| CBL13383.1 | 2.54e-213 | 4 | 499 | 13 | 514 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2BS9_A | 2.73e-147 | 8 | 486 | 6 | 484 | Nativecrystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_B Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_C Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_D Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_E Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_F Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_G Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_H Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 1W91_A | 2.20e-146 | 4 | 486 | 2 | 484 | crystalstructure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_B crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_C crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_D crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_E crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_F crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_G crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_H crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct] |
| 2BFG_A | 2.20e-146 | 8 | 486 | 6 | 484 | crystalstructure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_B crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_C crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_D crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_E crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_F crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_G crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_H crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus] |
| 1PX8_A | 3.25e-134 | 22 | 482 | 20 | 479 | Crystalstructure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1PX8_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_A Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_C Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_D Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum] |
| 6YYH_A | 2.51e-131 | 22 | 485 | 43 | 505 | Crystalstructure of beta-D-xylosidase from Dictyoglomus thermophilum in ligand-free form [Dictyoglomus thermophilum H-6-12],6YYH_B Crystal structure of beta-D-xylosidase from Dictyoglomus thermophilum in ligand-free form [Dictyoglomus thermophilum H-6-12],6YYI_A Crystal structure of beta-D-xylosidase from Dictyoglomus thermophilum bound to beta-D-xylopyranose [Dictyoglomus thermophilum H-6-12],6YYI_B Crystal structure of beta-D-xylosidase from Dictyoglomus thermophilum bound to beta-D-xylopyranose [Dictyoglomus thermophilum H-6-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9ZFM2 | 8.04e-144 | 8 | 486 | 6 | 485 | Beta-xylosidase OS=Geobacillus stearothermophilus OX=1422 GN=xynB PE=1 SV=1 |
| P36906 | 2.85e-132 | 22 | 482 | 20 | 479 | Beta-xylosidase OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=xynB PE=1 SV=1 |
| O30360 | 2.93e-129 | 22 | 482 | 20 | 479 | Beta-xylosidase OS=Thermoanaerobacterium saccharolyticum (strain DSM 8691 / JW/SL-YS485) OX=1094508 GN=xynB PE=3 SV=1 |
| P23552 | 2.99e-80 | 23 | 450 | 27 | 452 | Beta-xylosidase OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
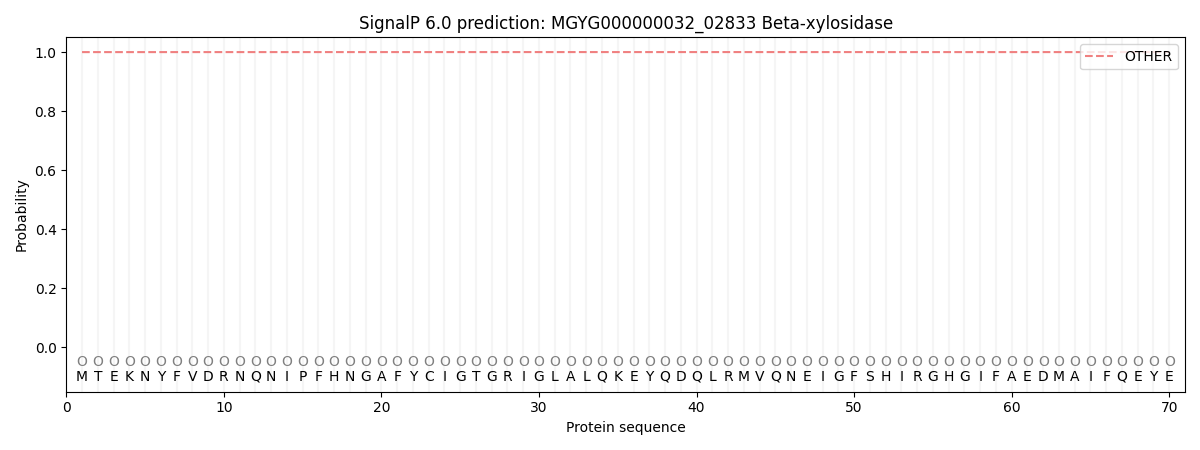
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000064 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
