You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000134_00705
You are here: Home > Sequence: MGYG000000134_00705
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
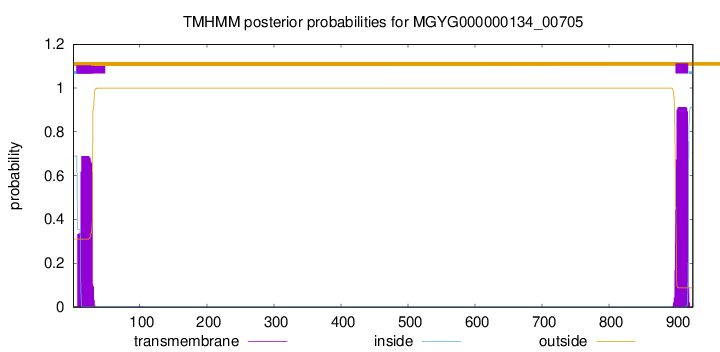
TMHMM annotations
Basic Information help
| Species | Longibaculum muris | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelatoclostridiaceae; Longibaculum; Longibaculum muris | |||||||||||
| CAZyme ID | MGYG000000134_00705 | |||||||||||
| CAZy Family | GH63 | |||||||||||
| CAZyme Description | Glucosidase YgjK | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 182907; End: 185684 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 21 | 785 | 5.5e-207 | 0.968421052631579 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10137 | PRK10137 | 0.0 | 40 | 783 | 31 | 782 | alpha-glucosidase; Provisional |
| pfam01204 | Trehalase | 1.21e-18 | 435 | 780 | 178 | 505 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| pfam03200 | Glyco_hydro_63 | 1.91e-12 | 583 | 774 | 252 | 486 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| COG3408 | GDB1 | 5.32e-11 | 330 | 757 | 259 | 578 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| PLN02567 | PLN02567 | 4.72e-10 | 435 | 773 | 203 | 533 | alpha,alpha-trehalase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QMW74875.1 | 0.0 | 1 | 924 | 1 | 913 |
| QPS11854.1 | 0.0 | 1 | 924 | 1 | 913 |
| QQV06747.1 | 0.0 | 1 | 924 | 1 | 913 |
| QQY26327.1 | 0.0 | 1 | 924 | 1 | 913 |
| QEK21330.1 | 0.0 | 6 | 799 | 4 | 804 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3W7S_A | 1.54e-155 | 41 | 783 | 5 | 755 | Escherichiacoli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7S_B Escherichia coli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7T_A Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7T_B Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7U_A Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12],3W7U_B Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12] |
| 3W7X_A | 8.50e-155 | 41 | 783 | 5 | 755 | Crystalstructure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],3W7X_B Crystal structure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],5CA3_A Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5CA3_B Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3W7W_A | 1.20e-154 | 41 | 783 | 5 | 755 | Crystalstructure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],3W7W_B Crystal structure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],5GW7_A Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5GW7_B Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3D3I_A | 3.71e-149 | 41 | 783 | 6 | 756 | Crystalstructural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12],3D3I_B Crystal structural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12] |
| 7PQQ_B | 4.61e-56 | 536 | 783 | 42 | 304 | ChainB, Anti-RON nanobody,Megabody 91,Glucosidase YgjK [Lama glama] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P42592 | 1.65e-154 | 41 | 783 | 28 | 778 | Glucosidase YgjK OS=Escherichia coli (strain K12) OX=83333 GN=ygjK PE=1 SV=1 |
| D8QTR2 | 7.40e-11 | 435 | 775 | 166 | 479 | Mannosylglycerate hydrolase MGH1 OS=Selaginella moellendorffii OX=88036 GN=MGH PE=1 SV=1 |
| D8T3S4 | 2.26e-10 | 435 | 775 | 166 | 479 | Mannosylglycerate hydrolase MGH2 OS=Selaginella moellendorffii OX=88036 GN=SELMODRAFT_447962 PE=3 SV=1 |
| P94250 | 7.47e-08 | 431 | 759 | 108 | 355 | Uncharacterized protein BB_0381 OS=Borreliella burgdorferi (strain ATCC 35210 / DSM 4680 / CIP 102532 / B31) OX=224326 GN=BB_0381 PE=4 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
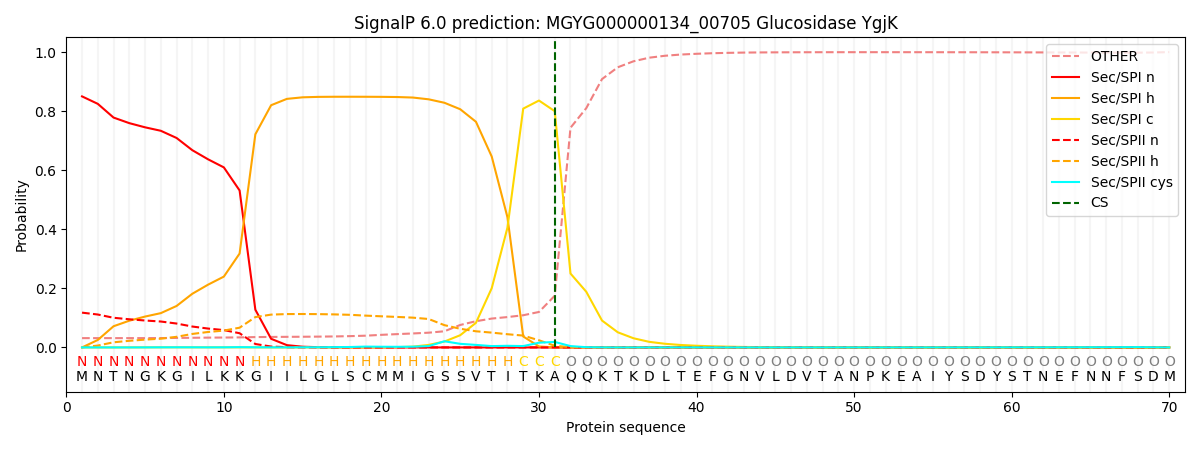
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.035199 | 0.840034 | 0.122933 | 0.001117 | 0.000373 | 0.000310 |