You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000138_01380
You are here: Home > Sequence: MGYG000000138_01380
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides johnsonii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides johnsonii | |||||||||||
| CAZyme ID | MGYG000000138_01380 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 368234; End: 369421 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 43 | 281 | 1.3e-61 | 0.995850622406639 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00413 | Glyco_hydrolase_16 | 1.93e-51 | 44 | 281 | 1 | 210 | glycosyl hydrolase family 16. The O-Glycosyl hydrolases are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycosyl hydrolase family 16. Family 16 includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
| cd08023 | GH16_laminarinase_like | 6.37e-34 | 42 | 281 | 1 | 235 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| cd02178 | GH16_beta_agarase | 2.18e-19 | 44 | 282 | 27 | 258 | Beta-agarase, member of glycosyl hydrolase family 16. Beta-agarase is a glycosyl hydrolase family 16 (GH16) member that hydrolyzes the internal beta-1,4-linkage of agarose, a hydrophilic polysaccharide found in the cell wall of Rhodophyceaea, marine red algae. Agarose is a linear chain of galactose units linked by alternating L-alpha-1,3- and D-beta-1,4-linkages that are additionally modified by a 3,6-anhydro-bridge. Agarose forms thermo-reversible gels that are widely used in the food industry or as a laboratory medium. While beta-agarases are also found in two other families derived from the sequence-based classification of glycosyl hydrolases (GH50, and GH86) the GH16 members are most abundant. This domain adopts a curved beta-sandwich conformation, with a tunnel-shaped active site cavity, referred to as a jellyroll fold. |
| pfam00722 | Glyco_hydro_16 | 5.28e-16 | 133 | 279 | 32 | 168 | Glycosyl hydrolases family 16. |
| cd02179 | GH16_beta_GRP | 3.75e-12 | 40 | 230 | 1 | 221 | beta-1,3-glucan recognition protein, member of glycosyl hydrolase family 16. Beta-GRP (beta-1,3-glucan recognition protein) is one of several pattern recognition receptors (PRRs), also referred to as biosensor proteins, that complexes with pathogen-associated beta-1,3-glucans and then transduces signals necessary for activation of an appropriate innate immune response. They are present in insects and lack all catalytic residues. This subgroup also contains related proteins of unknown function that still contain the active site. Their structures adopt a jelly roll fold with a deep active site channel harboring the catalytic residues, like those of other glycosyl hydrolase family 16 members. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT50278.1 | 3.03e-302 | 1 | 395 | 1 | 395 |
| AWS25493.1 | 4.24e-108 | 22 | 387 | 242 | 603 |
| ASY50995.1 | 4.24e-108 | 22 | 387 | 242 | 603 |
| QCJ40812.1 | 1.74e-80 | 29 | 375 | 114 | 482 |
| AIF44579.1 | 1.33e-78 | 29 | 370 | 111 | 474 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1UPS_A | 1.93e-36 | 24 | 317 | 16 | 321 | GlcNAc[alpha]1-4Galreleasing endo-[beta]-galactosidase from Clostridium perfringens [Clostridium perfringens],1UPS_B GlcNAc[alpha]1-4Gal releasing endo-[beta]-galactosidase from Clostridium perfringens [Clostridium perfringens] |
| 5WUT_A | 6.64e-19 | 36 | 282 | 3 | 235 | Crystalstructure of laminarinase from Flavobacterium sp. UMI-01 [Flavobacterium sp.],5WUT_B Crystal structure of laminarinase from Flavobacterium sp. UMI-01 [Flavobacterium sp.] |
| 6T2R_AAA | 9.95e-16 | 38 | 282 | 29 | 262 | ChainAAA, Beta-glycosidase [Bacteroides caccae] |
| 2VY0_A | 2.49e-15 | 33 | 281 | 8 | 259 | TheX-ray structure of endo-beta-1,3-glucanase from Pyrococcus furiosus [Pyrococcus furiosus],2VY0_B The X-ray structure of endo-beta-1,3-glucanase from Pyrococcus furiosus [Pyrococcus furiosus] |
| 6XOF_A | 3.76e-12 | 38 | 283 | 24 | 268 | ChainA, GH16 family protein [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23903 | 1.02e-09 | 37 | 283 | 422 | 681 | Glucan endo-1,3-beta-glucosidase A1 OS=Niallia circulans OX=1397 GN=glcA PE=1 SV=1 |
| Q84C00 | 6.86e-09 | 134 | 281 | 103 | 243 | Beta-glucanase OS=Acetivibrio thermocellus OX=1515 GN=licB PE=1 SV=1 |
| O33680 | 1.81e-08 | 124 | 281 | 308 | 457 | Endo-1,3-1,4-beta-glycanase ExsH OS=Rhizobium meliloti (strain 1021) OX=266834 GN=exsH PE=1 SV=1 |
| A3DBX3 | 3.89e-08 | 134 | 281 | 103 | 243 | Beta-glucanase OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=licB PE=1 SV=1 |
| Q9Z3Q2 | 4.10e-07 | 33 | 276 | 213 | 460 | Endo-1,3-1,4-beta-glycanase EglC OS=Rhizobium meliloti (strain 1021) OX=266834 GN=eglC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
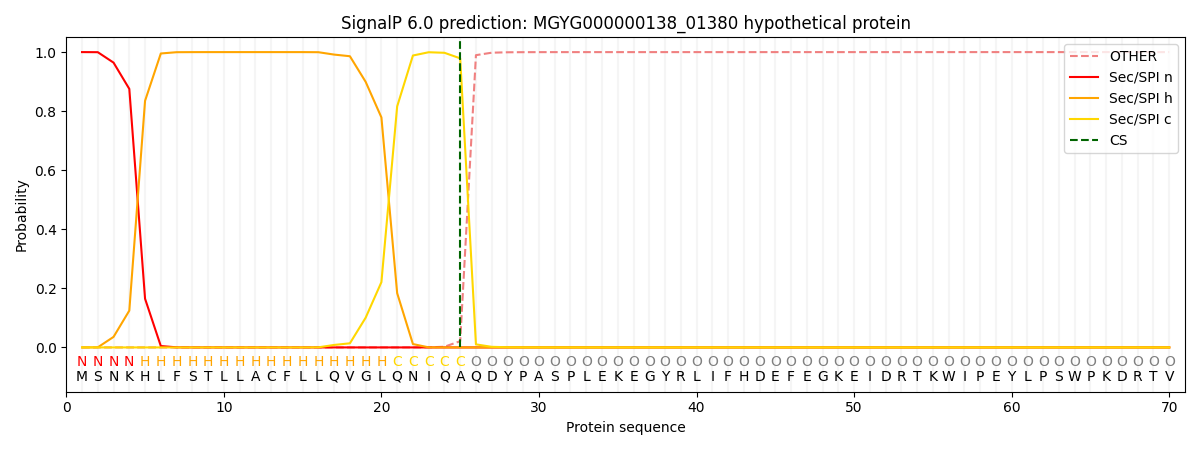
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000205 | 0.999198 | 0.000182 | 0.000146 | 0.000140 | 0.000128 |
