You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000138_03605
You are here: Home > Sequence: MGYG000000138_03605
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
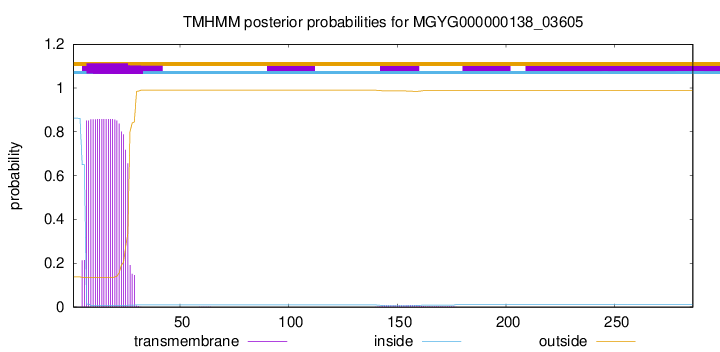
TMHMM annotations
Basic Information help
| Species | Parabacteroides johnsonii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides johnsonii | |||||||||||
| CAZyme ID | MGYG000000138_03605 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | Carbohydrate acetyl esterase/feruloyl esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3165; End: 4025 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 33 | 280 | 1.8e-62 | 0.9823788546255506 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2382 | Fes | 3.54e-39 | 33 | 278 | 75 | 294 | Enterochelin esterase or related enzyme [Inorganic ion transport and metabolism]. |
| pfam00756 | Esterase | 4.88e-36 | 33 | 278 | 1 | 245 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
| COG0627 | FrmB | 3.07e-32 | 40 | 283 | 35 | 312 | S-formylglutathione hydrolase FrmB [Defense mechanisms]. |
| COG2819 | YbbA | 3.44e-09 | 26 | 280 | 9 | 259 | Predicted hydrolase of the alpha/beta superfamily [General function prediction only]. |
| COG4099 | COG4099 | 7.73e-09 | 26 | 172 | 160 | 301 | Predicted peptidase [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABF40860.1 | 6.95e-38 | 31 | 272 | 149 | 374 |
| BCS34250.1 | 3.66e-35 | 31 | 272 | 110 | 335 |
| AKD58571.1 | 1.04e-34 | 31 | 283 | 145 | 380 |
| AMQ56447.1 | 1.12e-34 | 31 | 272 | 148 | 377 |
| QDU55105.1 | 6.90e-34 | 27 | 281 | 94 | 325 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5VOL_A | 2.06e-55 | 33 | 283 | 19 | 272 | Bacint_04212ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_B Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_C Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_D Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_E Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_F Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_G Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_H Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393] |
| 4RGY_A | 8.49e-27 | 31 | 279 | 9 | 261 | Structuraland functional analysis of a low-temperature-active alkaline esterase from South China Sea marine sediment microbial metagenomic library [uncultured bacterium FLS12],4RGY_B Structural and functional analysis of a low-temperature-active alkaline esterase from South China Sea marine sediment microbial metagenomic library [uncultured bacterium FLS12] |
| 1JJF_A | 1.11e-26 | 44 | 281 | 50 | 259 | ChainA, Endo-1,4-beta-xylanase Z [Acetivibrio thermocellus] |
| 1JT2_A | 3.02e-26 | 44 | 281 | 50 | 259 | ChainA, PROTEIN (ENDO-1,4-BETA-XYLANASE Z) [Acetivibrio thermocellus] |
| 6RZN_A | 3.53e-24 | 31 | 274 | 137 | 379 | Crystalstructure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium],6RZN_B Crystal structure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EXZ4 | 5.33e-27 | 44 | 276 | 449 | 665 | Carbohydrate acetyl esterase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe1-6A PE=1 SV=1 |
| P10478 | 1.93e-24 | 44 | 281 | 69 | 278 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
| P23553 | 3.07e-20 | 29 | 281 | 6 | 263 | Acetyl esterase OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynC PE=4 SV=1 |
| D5EY13 | 8.00e-19 | 27 | 275 | 501 | 720 | Endo-1,4-beta-xylanase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyn10D-fae1A PE=1 SV=1 |
| P9WM38 | 1.29e-16 | 31 | 263 | 179 | 425 | Esterase MT1326 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=MT1326 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
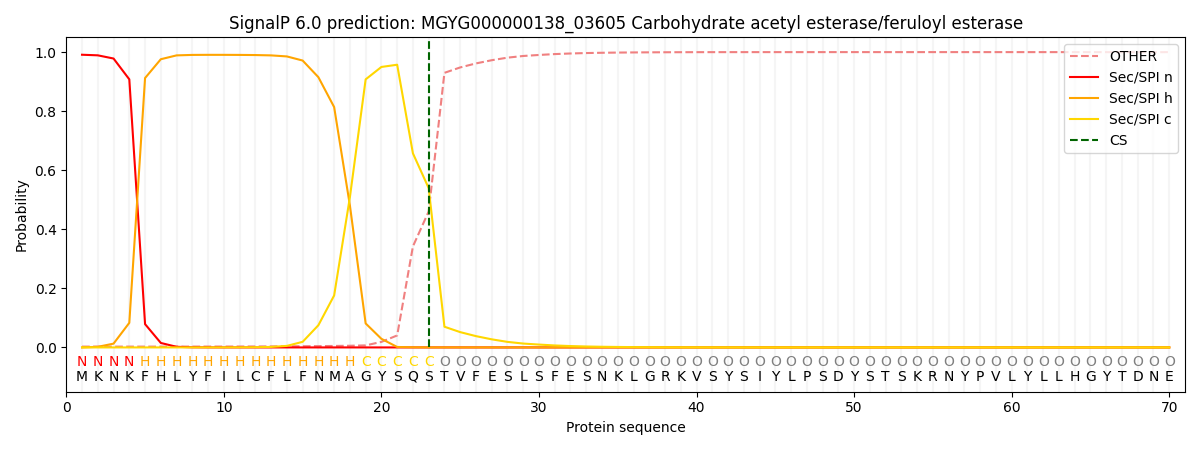
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.005161 | 0.986984 | 0.006965 | 0.000289 | 0.000272 | 0.000281 |