You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000219_03100
You are here: Home > Sequence: MGYG000000219_03100
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Eubacterium maltosivorans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Eubacteriales; Eubacteriaceae; Eubacterium; Eubacterium maltosivorans | |||||||||||
| CAZyme ID | MGYG000000219_03100 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 32683; End: 33540 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 2 | 231 | 1.9e-31 | 0.9217391304347826 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd04186 | GT_2_like_c | 6.01e-70 | 5 | 228 | 1 | 165 | Subfamily of Glycosyltransferase Family GT2 of unknown function. GT-2 includes diverse families of glycosyltransferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. These are enzymes that catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. Glycosyltransferases have been classified into more than 90 distinct sequence based families. |
| COG1216 | GT2 | 8.97e-56 | 1 | 281 | 3 | 270 | Glycosyltransferase, GT2 family [Carbohydrate transport and metabolism]. |
| cd02526 | GT2_RfbF_like | 5.35e-28 | 6 | 246 | 2 | 217 | RfbF is a putative dTDP-rhamnosyl transferase. Shigella flexneri RfbF protein is a putative dTDP-rhamnosyl transferase. dTDP rhamnosyl transferases of Shigella flexneri add rhamnose sugars to N-acetyl-glucosamine in the O-antigen tetrasaccharide repeat. Lipopolysaccharide O antigens are important virulence determinants for many bacteria. The variations of sugar composition, the sequence of the sugars and the linkages in the O antigen provide structural diversity of the O antigen. |
| pfam13641 | Glyco_tranf_2_3 | 1.40e-24 | 2 | 229 | 3 | 213 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| cd04185 | GT_2_like_b | 1.20e-22 | 6 | 246 | 2 | 194 | Subfamily of Glycosyltransferase Family GT2 of unknown function. GT-2 includes diverse families of glycosyltransferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. These are enzymes that catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. Glycosyltransferases have been classified into more than 90 distinct sequence based families. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCT71592.1 | 1.72e-207 | 1 | 285 | 1 | 285 |
| ALU13505.1 | 9.93e-207 | 1 | 285 | 1 | 285 |
| ADO38781.1 | 1.35e-204 | 1 | 285 | 1 | 285 |
| ARD65353.1 | 3.88e-204 | 1 | 285 | 1 | 285 |
| AFA47048.1 | 2.48e-152 | 1 | 285 | 8 | 292 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6P61_A | 5.38e-11 | 4 | 132 | 16 | 144 | Structureof a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_B Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_C Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_D Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197] |
| 5TZE_C | 6.50e-09 | 1 | 105 | 1 | 108 | Crystalstructure of S. aureus TarS in complex with UDP-GlcNAc [Staphylococcus aureus],5TZE_E Crystal structure of S. aureus TarS in complex with UDP-GlcNAc [Staphylococcus aureus],5TZI_C Crystal structure of S. aureus TarS 1-349 [Staphylococcus aureus],5TZJ_A Crystal structure of S. aureus TarS 1-349 in complex with UDP-GlcNAc [Staphylococcus aureus],5TZJ_C Crystal structure of S. aureus TarS 1-349 in complex with UDP-GlcNAc [Staphylococcus aureus],5TZK_C Crystal structure of S. aureus TarS 1-349 in complex with UDP [Staphylococcus aureus] |
| 5TZ8_A | 8.53e-09 | 1 | 105 | 1 | 108 | Crystalstructure of S. aureus TarS [Staphylococcus aureus],5TZ8_B Crystal structure of S. aureus TarS [Staphylococcus aureus],5TZ8_C Crystal structure of S. aureus TarS [Staphylococcus aureus] |
| 6YV7_B | 1.17e-07 | 4 | 223 | 45 | 242 | MannosyltransferasePcManGT from Pyrobaculum calidifontis [Pyrobaculum calidifontis JCM 11548],6YV8_B Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP and Mn2+ [Pyrobaculum calidifontis JCM 11548],6YV9_A Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP-Man and Mn2+ [Pyrobaculum calidifontis JCM 11548] |
| 6YV7_A | 1.18e-07 | 4 | 223 | 46 | 243 | MannosyltransferasePcManGT from Pyrobaculum calidifontis [Pyrobaculum calidifontis JCM 11548],6YV8_A Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP and Mn2+ [Pyrobaculum calidifontis JCM 11548],6YV9_B Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP-Man and Mn2+ [Pyrobaculum calidifontis JCM 11548] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P9WMY2 | 1.08e-19 | 3 | 285 | 5 | 296 | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=wbbL PE=3 SV=2 |
| P9WMY3 | 1.08e-19 | 3 | 285 | 5 | 296 | N-acetylglucosaminyl-diphospho-decaprenol L-rhamnosyltransferase OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=wbbL PE=1 SV=2 |
| D4GU63 | 6.26e-14 | 2 | 217 | 19 | 213 | Low-salt glycan biosynthesis hexosyltransferase Agl10 OS=Haloferax volcanii (strain ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2) OX=309800 GN=agl10 PE=3 SV=1 |
| P55465 | 3.20e-13 | 18 | 232 | 638 | 850 | Uncharacterized protein y4gI OS=Sinorhizobium fredii (strain NBRC 101917 / NGR234) OX=394 GN=NGR_a03550 PE=4 SV=1 |
| P36667 | 4.27e-12 | 67 | 228 | 61 | 220 | Rhamnosyltransferase WbbL OS=Escherichia coli (strain K12) OX=83333 GN=wbbL PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
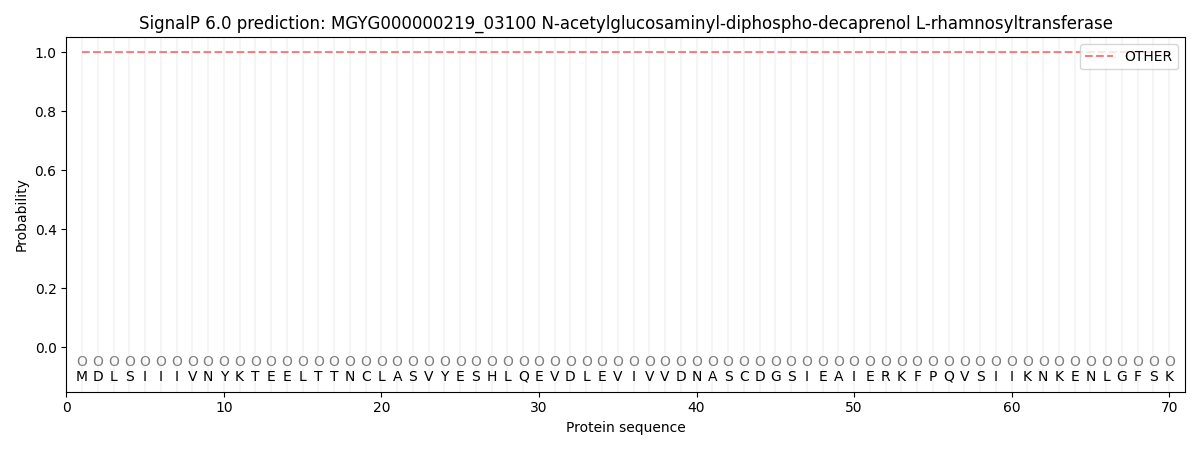
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000049 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
