You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000223_00065
You are here: Home > Sequence: MGYG000000223_00065
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | NSJ-61 sp003433845 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelotrichaceae; NSJ-61; NSJ-61 sp003433845 | |||||||||||
| CAZyme ID | MGYG000000223_00065 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 73660; End: 79989 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 496 | 733 | 6.8e-53 | 0.5339578454332553 |
| CBM32 | 1748 | 1873 | 2.6e-18 | 0.8629032258064516 |
| CBM32 | 1040 | 1146 | 1e-15 | 0.7983870967741935 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06596 | GH31_CPE1046 | 1.44e-154 | 281 | 704 | 1 | 334 | Clostridium CPE1046-like. CPE1046 is an uncharacterized Clostridium perfringens protein with a glycosyl hydrolase family 31 (GH31) domain. The domain architecture of CPE1046 and its orthologs includes a C-terminal fibronectin type 3 (FN3) domain and a coagulation factor 5/8 type C domain in addition to the GH31 domain. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| COG1501 | YicI | 7.97e-58 | 129 | 785 | 106 | 716 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| pfam01055 | Glyco_hydro_31 | 2.17e-50 | 490 | 733 | 211 | 442 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| cd08759 | Type_III_cohesin_like | 1.54e-36 | 1239 | 1409 | 1 | 167 | Cohesin domain, interaction partner of dockerin. Bacterial cohesin domains bind to a complementary protein domain named dockerin, and this interaction is required for the formation of the cellulosome, a cellulose-degrading complex. Two specific calcium-dependent interactions between cohesin and dockerin appear to be essential for cellulosome assembly, type I and type II. This subfamily represents type III cohesins and closely related domains. |
| cd06589 | GH31 | 3.28e-34 | 365 | 616 | 25 | 265 | glycosyl hydrolase family 31 (GH31). GH31 enzymes occur in prokaryotes, eukaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. In most cases, the pyranose moiety recognized in subsite -1 of the substrate binding site is an alpha-D-glucose, though some GH31 family members show a preference for alpha-D-xylose. Several GH31 enzymes can accommodate both glucose and xylose and different levels of discrimination between the two have been observed. Most characterized GH31 enzymes are alpha-glucosidases. In mammals, GH31 members with alpha-glucosidase activity are implicated in at least three distinct biological processes. The lysosomal acid alpha-glucosidase (GAA) is essential for glycogen degradation and a deficiency or malfunction of this enzyme causes glycogen storage disease II, also known as Pompe disease. In the endoplasmic reticulum, alpha-glucosidase II catalyzes the second step in the N-linked oligosaccharide processing pathway that constitutes part of the quality control system for glycoprotein folding and maturation. The intestinal enzymes sucrase-isomaltase (SI) and maltase-glucoamylase (MGAM) play key roles in the final stage of carbohydrate digestion, making alpha-glucosidase inhibitors useful in the treatment of type 2 diabetes. GH31 alpha-glycosidases are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNM10857.1 | 0.0 | 1 | 2109 | 1 | 2109 |
| BCT46261.1 | 0.0 | 1 | 2024 | 1 | 2051 |
| BBK61154.1 | 0.0 | 6 | 2018 | 1 | 2074 |
| QWT17625.1 | 0.0 | 42 | 2014 | 45 | 2102 |
| BBK23937.1 | 0.0 | 42 | 1880 | 55 | 1851 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6M76_A | 5.01e-251 | 39 | 1024 | 32 | 963 | GH31alpha-N-acetylgalactosaminidase from Enterococcus faecalis [Enterococcus faecalis ATCC 10100],6M77_A GH31 alpha-N-acetylgalactosaminidase from Enterococcus faecalis in complex with N-acetylgalactosamine [Enterococcus faecalis ATCC 10100] |
| 7F7R_A | 2.69e-250 | 39 | 1024 | 32 | 963 | ChainA, GH31 alpha-N-acetylgalactosaminidase [Enterococcus faecalis ATCC 10100] |
| 7F7Q_A | 7.38e-250 | 39 | 1024 | 32 | 963 | ChainA, GH31 alpha-N-acetylgalactosaminidase [Enterococcus faecalis ATCC 10100] |
| 6JR6_A | 7.15e-21 | 177 | 896 | 154 | 835 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR7_A Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101] |
| 6JR8_A | 7.15e-21 | 177 | 896 | 154 | 835 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9F234 | 7.84e-22 | 511 | 759 | 462 | 692 | Alpha-glucosidase 2 OS=Bacillus thermoamyloliquefaciens OX=1425 PE=3 SV=1 |
| P38138 | 3.96e-17 | 504 | 747 | 585 | 823 | Glucosidase 2 subunit alpha OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=ROT2 PE=1 SV=1 |
| Q8BVW0 | 1.12e-16 | 506 | 735 | 544 | 765 | Neutral alpha-glucosidase C OS=Mus musculus OX=10090 GN=Ganc PE=1 SV=2 |
| Q09901 | 2.08e-16 | 494 | 796 | 641 | 923 | Uncharacterized family 31 glucosidase C30D11.01c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC30D11.01c PE=3 SV=2 |
| Q9URX4 | 3.57e-16 | 502 | 784 | 636 | 908 | Uncharacterized family 31 glucosidase C1039.11c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC1039.11c PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
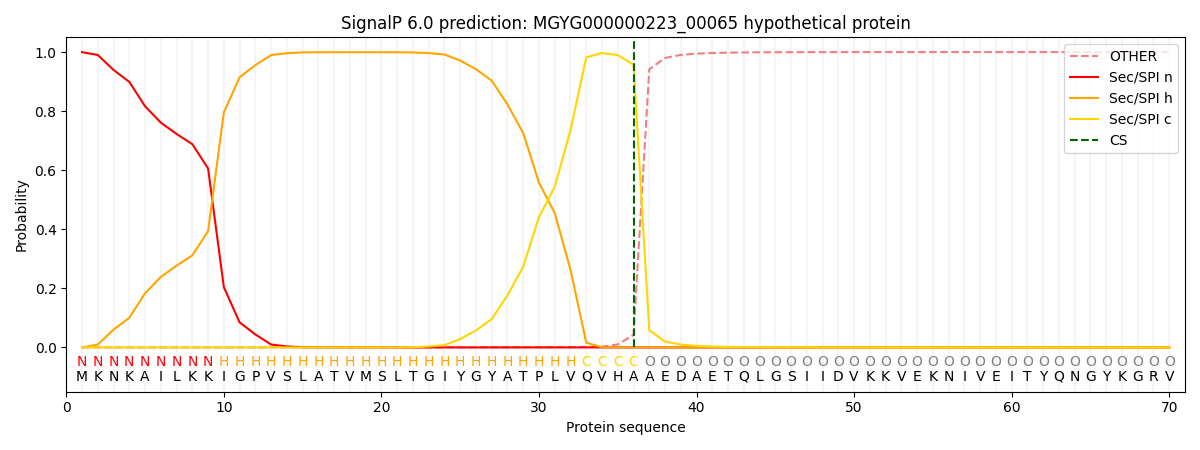
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000659 | 0.998509 | 0.000196 | 0.000248 | 0.000183 | 0.000164 |
