You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000234_00638
You are here: Home > Sequence: MGYG000000234_00638
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Vagococcus teuberi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Vagococcaceae; Vagococcus; Vagococcus teuberi | |||||||||||
| CAZyme ID | MGYG000000234_00638 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 654132; End: 656255 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK08581 | PRK08581 | 1.02e-70 | 210 | 552 | 132 | 472 | amidase domain-containing protein. |
| COG1705 | FlgJ | 1.03e-57 | 373 | 554 | 22 | 190 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK06347 | PRK06347 | 2.71e-52 | 291 | 558 | 51 | 309 | 1,4-beta-N-acetylmuramoylhydrolase. |
| COG0739 | NlpD | 4.10e-36 | 473 | 704 | 27 | 263 | Murein DD-endopeptidase MepM and murein hydrolase activator NlpD, contain LysM domain [Cell wall/membrane/envelope biogenesis]. |
| cd12797 | M23_peptidase | 1.87e-35 | 606 | 690 | 1 | 85 | M23 family metallopeptidase, also known as beta-lytic metallopeptidase, and similar proteins. This model describes the metallopeptidase M23 family, which includes beta-lytic metallopeptidase and lysostaphin. Members of this family are zinc endopeptidases that lyse bacterial cell wall peptidoglycans; they cleave either the N-acylmuramoyl-Ala bond between the cell wall peptidoglycan and the cross-linking peptide (e.g. beta-lytic endopeptidase) or a bond within the cross-linking peptide (e.g. stapholysin, and lysostaphin). Beta-lytic metallopeptidase, formerly known as beta-lytic protease, has a preference for cleavage of Gly-X bonds and favors hydrophobic or apolar residues on either side. It inhibits growth of sensitive organisms and may potentially serve as an antimicrobial agent. Lysostaphin, produced by Staphylococcus genus, cleaves pentaglycine cross-bridges of cell wall peptidoglycan, acting as autolysins to maintain cell wall metabolism or as toxins and weapons against competing strains. Staphylolysin (also known as LasA) is implicated in a range of processes related to Pseudomonas virulence, including stimulating shedding of the ectodomain of cell surface heparan sulphate proteoglycan syndecan-1, and elastin degradation in connective tissue. Its active site is less constricted and contains a five-coordinate zinc ion with trigonal bipyramidal geometry and two metal-bound water molecules, possibly contributing to its activity against a wider range of substrates than those used by related lytic enzymes, consistent with its multiple roles in Pseudomonas virulence. The family includes members that do not appear to have the conserved zinc-binding site and might be lipoproteins lacking proteolytic activity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APB31703.1 | 9.73e-212 | 389 | 707 | 1 | 313 |
| QNN74568.1 | 5.23e-120 | 383 | 707 | 138 | 455 |
| QIL47333.1 | 5.12e-119 | 385 | 707 | 166 | 529 |
| QNN73822.1 | 2.28e-118 | 383 | 706 | 168 | 532 |
| QCT93447.1 | 2.42e-118 | 393 | 707 | 227 | 535 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5T1Q_A | 6.67e-43 | 359 | 552 | 26 | 212 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 3FI7_A | 8.83e-38 | 394 | 552 | 31 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 6UE4_A | 8.08e-21 | 553 | 688 | 203 | 348 | ShyAEndopeptidase from Vibrio cholerae (Closed form) [Vibrio cholerae O1 biovar El Tor str. N16961],6UE4_B ShyA Endopeptidase from Vibrio cholerae (Closed form) [Vibrio cholerae O1 biovar El Tor str. N16961] |
| 6U2A_A | 8.34e-21 | 553 | 688 | 200 | 345 | ShyAendopeptidase from Vibrio cholera (open form) [Vibrio cholerae] |
| 3SLU_A | 3.90e-17 | 594 | 688 | 228 | 327 | Crystalstructure of NMB0315 [Neisseria meningitidis ATCC 13091],3SLU_B Crystal structure of NMB0315 [Neisseria meningitidis ATCC 13091] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q2G222 | 4.83e-41 | 359 | 552 | 286 | 472 | N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=SAOUHSC_02979 PE=1 SV=1 |
| P39046 | 1.60e-28 | 383 | 565 | 52 | 229 | Muramidase-2 OS=Enterococcus hirae (strain ATCC 9790 / DSM 20160 / JCM 8729 / LMG 6399 / NBRC 3181 / NCIMB 6459 / NCDO 1258 / NCTC 12367 / WDCM 00089 / R) OX=768486 GN=EHR_05900 PE=1 SV=1 |
| P37710 | 4.80e-28 | 388 | 552 | 175 | 333 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| Q9CIT4 | 1.60e-26 | 383 | 565 | 52 | 227 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
| A2RHZ5 | 6.84e-26 | 383 | 558 | 52 | 220 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
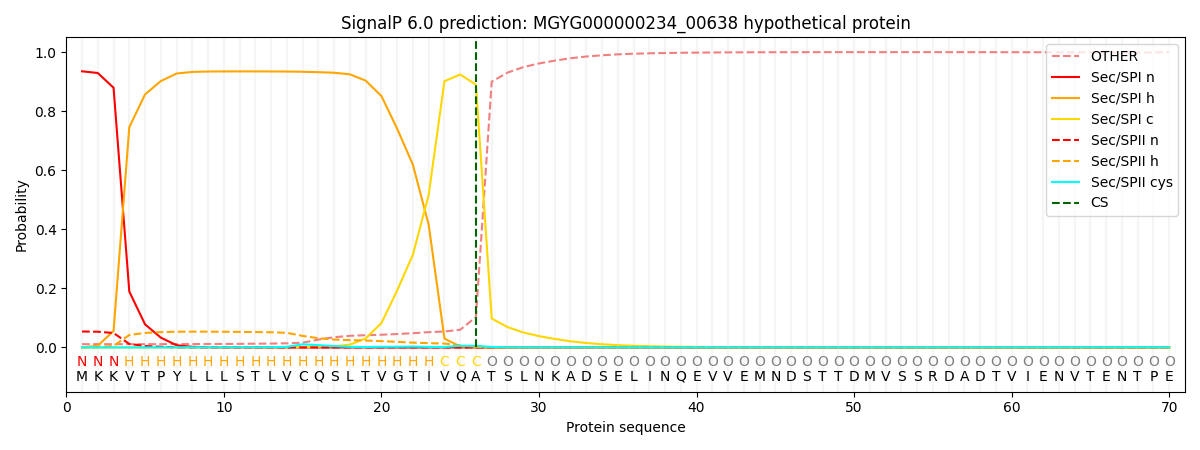
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.015670 | 0.925751 | 0.057556 | 0.000364 | 0.000306 | 0.000312 |
