You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000234_01770
You are here: Home > Sequence: MGYG000000234_01770
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Vagococcus teuberi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Vagococcaceae; Vagococcus; Vagococcus teuberi | |||||||||||
| CAZyme ID | MGYG000000234_01770 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 49387; End: 51981 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH73 | 318 | 459 | 3e-28 | 0.984375 |
| CBM50 | 498 | 539 | 1.8e-16 | 0.95 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK06347 | PRK06347 | 1.02e-106 | 213 | 748 | 51 | 591 | 1,4-beta-N-acetylmuramoylhydrolase. |
| COG1705 | FlgJ | 1.51e-58 | 310 | 466 | 43 | 191 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK08581 | PRK08581 | 4.29e-52 | 1 | 464 | 1 | 473 | amidase domain-containing protein. |
| smart00047 | LYZ2 | 3.18e-33 | 311 | 463 | 9 | 147 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| pfam01832 | Glucosaminidase | 2.50e-24 | 318 | 401 | 1 | 76 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APB30413.1 | 0.0 | 334 | 864 | 1 | 531 |
| QIL47903.1 | 5.44e-195 | 1 | 863 | 1 | 1001 |
| QNN73119.1 | 1.31e-182 | 1 | 863 | 1 | 978 |
| QOG33096.1 | 5.38e-131 | 294 | 863 | 158 | 736 |
| QCA28964.1 | 7.82e-131 | 291 | 864 | 69 | 667 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FI7_A | 8.51e-44 | 306 | 463 | 24 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 5T1Q_A | 1.72e-35 | 313 | 464 | 63 | 213 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 2MKX_A | 3.01e-10 | 492 | 543 | 1 | 52 | Solutionstructure of LysM the peptidoglycan binding domain of autolysin AtlA from Enterococcus faecalis [Enterococcus faecalis V583] |
| 4UZ2_A | 1.84e-07 | 497 | 545 | 4 | 52 | Crystalstructure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_B Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_C Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_D Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ3_A Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8],4UZ3_B Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8],4UZ3_C Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8] |
| 4XCM_A | 2.17e-06 | 497 | 545 | 4 | 52 | Crystalstructure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4XCM_B Crystal structure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P37710 | 5.88e-131 | 294 | 863 | 158 | 736 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| P39046 | 2.00e-106 | 311 | 863 | 63 | 664 | Muramidase-2 OS=Enterococcus hirae (strain ATCC 9790 / DSM 20160 / JCM 8729 / LMG 6399 / NBRC 3181 / NCIMB 6459 / NCDO 1258 / NCTC 12367 / WDCM 00089 / R) OX=768486 GN=EHR_05900 PE=1 SV=1 |
| Q9CIT4 | 1.46e-77 | 313 | 687 | 65 | 438 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
| A2RHZ5 | 2.67e-76 | 313 | 687 | 65 | 436 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
| P0C2T5 | 2.68e-75 | 313 | 687 | 65 | 436 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris OX=1359 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
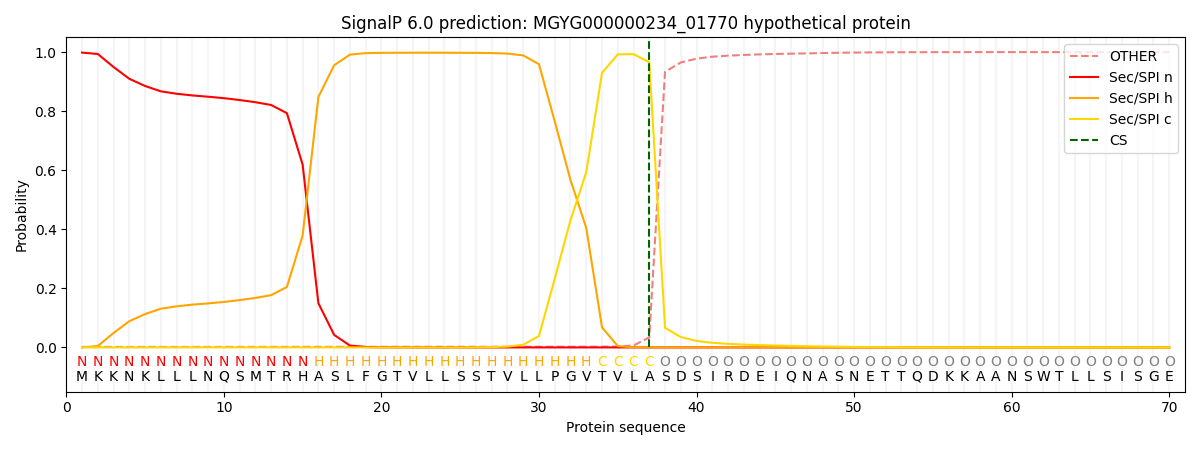
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.003116 | 0.995741 | 0.000277 | 0.000336 | 0.000264 | 0.000245 |
