You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000441_00681
You are here: Home > Sequence: MGYG000000441_00681
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
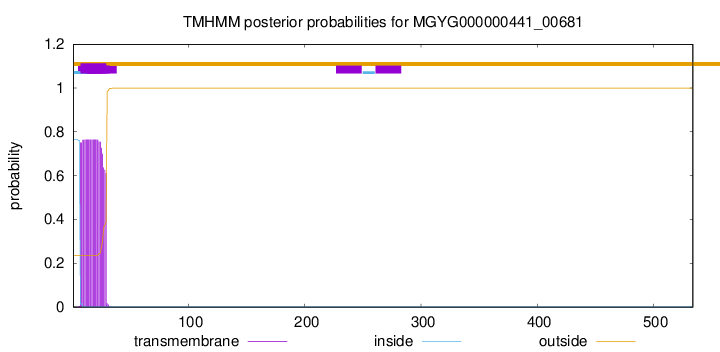
TMHMM annotations
Basic Information help
| Species | CAG-312 sp900545705 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; CAG-312 sp900545705 | |||||||||||
| CAZyme ID | MGYG000000441_00681 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 51321; End: 52925 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 142 | 531 | 6.1e-85 | 0.9110070257611241 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06592 | GH31_NET37 | 4.68e-175 | 137 | 504 | 1 | 364 | glucosidase NET37. NET37 (also known as KIAA1161) is a human lamina-associated nuclear envelope transmembrane protein. A member of the glycosyl hydrolase family 31 (GH31) , it has been shown to be required for myogenic differentiation of C2C12 cells. Related proteins are found in eukaryotes and prokaryotes. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| pfam01055 | Glyco_hydro_31 | 1.34e-58 | 151 | 530 | 40 | 435 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| COG1501 | YicI | 3.11e-57 | 33 | 532 | 151 | 665 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| cd06597 | GH31_transferase_CtsY | 2.23e-30 | 144 | 436 | 14 | 324 | CtsY (cyclic tetrasaccharide-synthesizing enzyme Y)-like. CtsY is a bacterial 3-alpha-isomaltosyltransferase, first identified in Arthrobacter globiformis, that produces cyclic tetrasaccharides together with a closely related enzyme CtsZ. CtsY and CtsZ both have a glycosyl hydrolase family 31 (GH31) catalytic domain; CtsZ belongs to a different subfamily. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| PRK10426 | PRK10426 | 1.36e-27 | 69 | 530 | 140 | 616 | alpha-glucosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT50281.1 | 5.31e-257 | 3 | 534 | 1 | 531 |
| QCT77881.1 | 1.32e-238 | 2 | 533 | 22 | 549 |
| CAH07048.1 | 1.32e-238 | 2 | 533 | 22 | 549 |
| CUA17940.1 | 1.32e-238 | 2 | 533 | 22 | 549 |
| QCQ41002.1 | 1.32e-238 | 2 | 533 | 22 | 549 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2F2H_A | 1.48e-28 | 141 | 524 | 269 | 661 | Structureof the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_B Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_C Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_D Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_E Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_F Structure of the YicI thiosugar Michaelis complex [Escherichia coli] |
| 1XSI_A | 1.49e-28 | 141 | 524 | 269 | 661 | Structureof a Family 31 alpha glycosidase [Escherichia coli],1XSI_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_A Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSK_A Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_B Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_C Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_D Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_E Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_F Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli] |
| 1WE5_A | 6.32e-25 | 141 | 524 | 269 | 661 | CrystalStructure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_B Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_C Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_D Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_E Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli],1WE5_F Crystal Structure of Alpha-Xylosidase from Escherichia coli [Escherichia coli] |
| 6JR6_A | 3.41e-20 | 165 | 532 | 292 | 683 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR6_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A [Flavobacterium johnsoniae UW101],6JR7_A Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101],6JR7_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, complexed with glucose [Flavobacterium johnsoniae UW101] |
| 6JR8_A | 1.86e-19 | 165 | 532 | 292 | 683 | Flavobacteriumjohnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_B Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_C Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101],6JR8_D Flavobacterium johnsoniae GH31 dextranase, FjDex31A, mutant D412A complexed with isomaltotriose [Flavobacterium johnsoniae UW101] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q6NSJ0 | 4.79e-61 | 106 | 530 | 283 | 710 | Myogenesis-regulating glycosidase OS=Homo sapiens OX=9606 GN=MYORG PE=1 SV=2 |
| Q69ZQ1 | 1.75e-59 | 106 | 525 | 282 | 705 | Myogenesis-regulating glycosidase OS=Mus musculus OX=10090 GN=Myorg PE=1 SV=2 |
| P31434 | 8.09e-28 | 141 | 524 | 269 | 661 | Alpha-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yicI PE=1 SV=2 |
| P96793 | 1.08e-25 | 35 | 513 | 158 | 647 | Alpha-xylosidase XylQ OS=Lactiplantibacillus pentosus OX=1589 GN=xylQ PE=1 SV=1 |
| Q9F234 | 7.85e-21 | 151 | 532 | 272 | 669 | Alpha-glucosidase 2 OS=Bacillus thermoamyloliquefaciens OX=1425 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000237 | 0.999078 | 0.000169 | 0.000170 | 0.000168 | 0.000146 |