You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000508_00834
You are here: Home > Sequence: MGYG000000508_00834
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-724 sp003524145 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; CAG-272; CAG-724; CAG-724 sp003524145 | |||||||||||
| CAZyme ID | MGYG000000508_00834 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | Elongation factor 4 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7257; End: 9791 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05306 | infB | 0.0 | 143 | 844 | 52 | 745 | translation initiation factor IF-2; Validated |
| TIGR00487 | IF-2 | 0.0 | 266 | 844 | 5 | 584 | translation initiation factor IF-2. This model discriminates eubacterial (and mitochondrial) translation initiation factor 2 (IF-2), encoded by the infB gene in bacteria, from similar proteins in the Archaea and Eukaryotes. In the bacteria and in organelles, the initiator tRNA is charged with N-formyl-Met instead of Met. This translation factor acts in delivering the initator tRNA to the ribosome. It is one of a number of GTP-binding translation factors recognized by the pfam model GTP_EFTU. [Protein synthesis, Translation factors] |
| CHL00189 | infB | 0.0 | 219 | 844 | 118 | 740 | translation initiation factor 2; Provisional |
| COG0532 | InfB | 0.0 | 345 | 844 | 1 | 505 | Translation initiation factor IF-2, a GTPase [Translation, ribosomal structure and biogenesis]. |
| cd01887 | IF2_eIF5B | 1.28e-99 | 350 | 512 | 1 | 168 | Initiation Factor 2 (IF2)/ eukaryotic Initiation Factor 5B (eIF5B) family. IF2/eIF5B contribute to ribosomal subunit joining and function as GTPases that are maximally activated by the presence of both ribosomal subunits. As seen in other GTPases, IF2/IF5B undergoes conformational changes between its GTP- and GDP-bound states. Eukaryotic IF2/eIF5Bs possess three characteristic segments, including a divergent N-terminal region followed by conserved central and C-terminal segments. This core region is conserved among all known eukaryotic and archaeal IF2/eIF5Bs and eubacterial IF2s. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGL64345.2 | 2.29e-202 | 287 | 842 | 297 | 855 |
| CAE6204650.1 | 9.01e-11 | 354 | 555 | 752 | 970 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3JCJ_f | 3.43e-196 | 267 | 844 | 310 | 887 | Structuresof ribosome-bound initiation factor 2 reveal the mechanism of subunit association [Escherichia coli],3JCN_b Structures of ribosome-bound initiation factor 2 reveal the mechanism of subunit association: Initiation Complex I [Escherichia coli],5ME0_W Chain W, Translation initiation factor IF-2 [Escherichia coli K-12],5ME1_W Structure of the 30S Pre-Initiation Complex 2 (30S IC-2) Stalled by GE81112 [Escherichia coli K-12] |
| 6O7K_f | 2.43e-185 | 348 | 844 | 9 | 506 | 30Sinitiation complex [Escherichia coli],6O9K_z 70S initiation complex [Escherichia coli] |
| 1ZO1_I | 7.35e-185 | 348 | 844 | 3 | 500 | IF2,IF1, and tRNA fitted to cryo-EM data OF E. COLI 70S initiation complex [Escherichia coli] |
| 5LMV_a | 2.18e-156 | 316 | 842 | 39 | 569 | Structureof bacterial 30S-IF1-IF2-IF3-mRNA-tRNA translation pre-initiation complex(state-III) [Thermus thermophilus HB8] |
| 3J4J_A | 8.09e-156 | 316 | 842 | 39 | 569 | Modelof full-length T. thermophilus Translation Initiation Factor 2 refined against its cryo-EM density from a 30S Initiation Complex map [Thermus thermophilus HB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DE44 | 8.90e-247 | 267 | 842 | 456 | 1033 | Translation initiation factor IF-2 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=infB PE=3 SV=1 |
| A4XL70 | 9.04e-246 | 267 | 842 | 281 | 856 | Translation initiation factor IF-2 OS=Caldicellulosiruptor saccharolyticus (strain ATCC 43494 / DSM 8903 / Tp8T 6331) OX=351627 GN=infB PE=3 SV=1 |
| Q9KA77 | 5.15e-244 | 267 | 844 | 151 | 729 | Translation initiation factor IF-2 OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=infB PE=3 SV=1 |
| Q3AB98 | 8.39e-244 | 267 | 844 | 246 | 824 | Translation initiation factor IF-2 OS=Carboxydothermus hydrogenoformans (strain ATCC BAA-161 / DSM 6008 / Z-2901) OX=246194 GN=infB PE=3 SV=1 |
| B2TJ55 | 9.44e-244 | 267 | 844 | 107 | 684 | Translation initiation factor IF-2 OS=Clostridium botulinum (strain Eklund 17B / Type B) OX=935198 GN=infB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
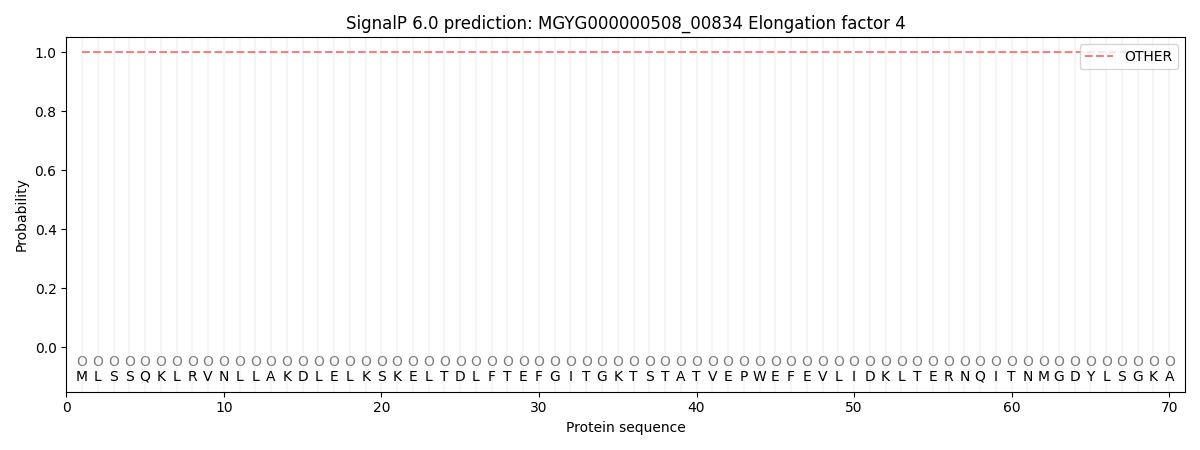
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000053 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
