You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000952_00777
You are here: Home > Sequence: MGYG000000952_00777
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA4855 sp900540365 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; RFN20; CAG-826; UBA4855; UBA4855 sp900540365 | |||||||||||
| CAZyme ID | MGYG000000952_00777 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5340; End: 8366 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 465 | 893 | 2e-76 | 0.49069148936170215 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 3.82e-32 | 470 | 877 | 13 | 426 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| cd17325 | MFS_MdtG_SLC18_like | 8.03e-27 | 35 | 389 | 21 | 351 | bacterial MdtG-like and eukaryotic solute carrier 18 (SLC18) family of the Major Facilitator Superfamily of transporters. This family is composed of eukaryotic solute carrier 18 (SLC18) family transporters and related bacterial multidrug resistance (MDR) transporters including several proteins from Escherichia coli such as multidrug resistance protein MdtG, from Bacillus subtilis such as multidrug resistance proteins 1 (Bmr1) and 2 (Bmr2), and from Staphylococcus aureus such as quinolone resistance protein NorA. The family also includes Escherichia coli arabinose efflux transporters YfcJ and YhhS. MDR transporters are drug/H+ antiporters (DHA) that mediate the efflux of a variety of drugs and toxic compounds, and confer resistance to these compounds. The SLC18 transporter family includes vesicular monoamine transporters (VAT1 and VAT2), vesicular acetylcholine transporter (VAChT), and SLC18B1, which is proposed to be a vesicular polyamine transporter (VPAT). The MdtG/SLC18 family belongs to the Major Facilitator Superfamily (MFS) of membrane transport proteins, which are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. |
| cd06174 | MFS | 1.00e-26 | 27 | 395 | 13 | 362 | Major Facilitator Superfamily. The Major Facilitator Superfamily (MFS) is a large and diverse group of secondary transporters that includes uniporters, symporters, and antiporters. MFS proteins facilitate the transport across cytoplasmic or internal membranes of a variety of substrates including ions, sugar phosphates, drugs, neurotransmitters, nucleosides, amino acids, and peptides. They do so using the electrochemical potential of the transported substrates. Uniporters transport a single substrate, while symporters and antiporters transport two substrates in the same or in opposite directions, respectively, across membranes. MFS proteins are typically 400 to 600 amino acids in length, and the majority contain 12 transmembrane alpha helices (TMs) connected by hydrophilic loops. The N- and C-terminal halves of these proteins display weak similarity and may be the result of a gene duplication/fusion event. Based on kinetic studies and the structures of a few bacterial superfamily members, GlpT (glycerol-3-phosphate transporter), LacY (lactose permease), and EmrD (multidrug transporter), MFS proteins are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. Bacterial members function primarily for nutrient uptake, and as drug-efflux pumps to confer antibiotic resistance. Some MFS proteins have medical significance in humans such as the glucose transporter Glut4, which is impaired in type II diabetes, and glucose-6-phosphate transporter (G6PT), which causes glycogen storage disease when mutated. |
| pfam07690 | MFS_1 | 9.93e-26 | 46 | 379 | 33 | 346 | Major Facilitator Superfamily. |
| PRK10150 | PRK10150 | 2.75e-22 | 519 | 920 | 69 | 510 | beta-D-glucuronidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABX41098.1 | 1.84e-127 | 455 | 1003 | 16 | 615 |
| VEU80886.1 | 2.62e-123 | 456 | 1007 | 25 | 580 |
| QYX26866.1 | 3.50e-122 | 456 | 1006 | 18 | 578 |
| QIK56880.1 | 3.36e-121 | 451 | 1003 | 7 | 575 |
| CUH91999.1 | 6.72e-121 | 452 | 999 | 32 | 601 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7SF2_A | 2.76e-78 | 456 | 1006 | 26 | 582 | ChainA, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_E Chain E, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_F Chain F, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838] |
| 6U7J_A | 6.02e-22 | 500 | 920 | 53 | 508 | UnculturedClostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_B Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_C Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_D Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.] |
| 4JKM_A | 8.24e-22 | 459 | 920 | 6 | 510 | CrystalStructure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],4JKM_B Crystal Structure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],6CXS_A Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13],6CXS_B Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13] |
| 6XXW_A | 3.29e-21 | 459 | 881 | 23 | 439 | Structureof beta-D-Glucuronidase for Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
| 6S6Z_A | 4.71e-20 | 458 | 879 | 33 | 462 | Structureof beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_B Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_C Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_D Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_E Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_F Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_G Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8],6S6Z_H Structure of beta-Galactosidase from Thermotoga maritima [Thermotoga maritima MSB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q56307 | 2.58e-19 | 458 | 879 | 34 | 463 | Beta-galactosidase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=lacZ PE=1 SV=2 |
| T2KPJ7 | 1.08e-18 | 519 | 857 | 107 | 430 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| T2KM09 | 4.75e-13 | 519 | 885 | 110 | 459 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22050 PE=2 SV=2 |
| P05804 | 1.28e-12 | 517 | 857 | 67 | 413 | Beta-glucuronidase OS=Escherichia coli (strain K12) OX=83333 GN=uidA PE=1 SV=2 |
| P77989 | 1.38e-11 | 508 | 924 | 46 | 459 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
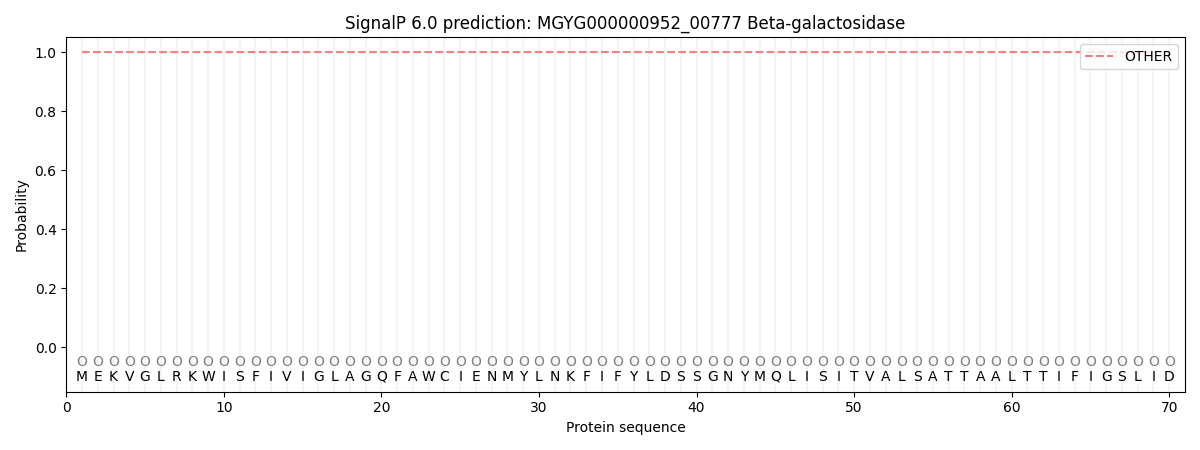
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
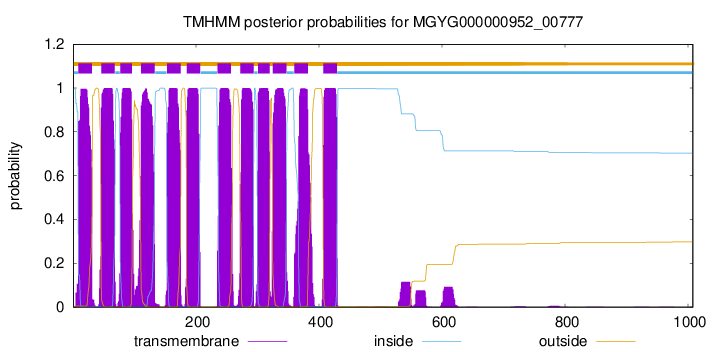
| start | end |
|---|---|
| 9 | 31 |
| 46 | 68 |
| 77 | 96 |
| 111 | 133 |
| 153 | 175 |
| 185 | 207 |
| 235 | 257 |
| 272 | 294 |
| 301 | 320 |
| 325 | 347 |
| 360 | 382 |
| 407 | 429 |
