You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001323_00575
You are here: Home > Sequence: MGYG000001323_00575
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Providencia rettgeri_D | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Providencia; Providencia rettgeri_D | |||||||||||
| CAZyme ID | MGYG000001323_00575 | |||||||||||
| CAZy Family | GT41 | |||||||||||
| CAZyme Description | UDP-glucose:protein N-beta-glucosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 107881; End: 109788 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT41 | 281 | 621 | 3.2e-37 | 0.4326241134751773 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam18071 | HMW1C_N | 1.77e-29 | 15 | 154 | 2 | 143 | HMW1C N-terminal. This is the N-terminal domain found in Actinobacillus pleuropneumoniae HMW1C (ApHMW1C). HMW1 adhesin is an N-linked glycoprotein that mediates adherence to respiratory epithelium through N-glycosylation of protein acceptor sites an O-glycosylation of sugar acceptor sites. The N-terminal domain forms an all alpha domain (AAD) when combined with the domain spanning from residue 154 to residue 245. The AAD interacts extensively with the C-terminal GT-B fold in order to create unique groove with potential to accommodate the acceptor protein. |
| COG3914 | Spy | 6.30e-13 | 363 | 627 | 356 | 613 | Predicted O-linked N-acetylglucosamine transferase, SPINDLY family [Posttranslational modification, protein turnover, chaperones]. |
| cd03801 | GT4_PimA-like | 0.007 | 272 | 596 | 2 | 340 | phosphatidyl-myo-inositol mannosyltransferase. This family is most closely related to the GT4 family of glycosyltransferases and named after PimA in Propionibacterium freudenreichii, which is involved in the biosynthesis of phosphatidyl-myo-inositol mannosides (PIM) which are early precursors in the biosynthesis of lipomannans (LM) and lipoarabinomannans (LAM), and catalyzes the addition of a mannosyl residue from GDP-D-mannose (GDP-Man) to the position 2 of the carrier lipid phosphatidyl-myo-inositol (PI) to generate a phosphatidyl-myo-inositol bearing an alpha-1,2-linked mannose residue (PIM1). Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CUR95075.1 | 7.03e-259 | 15 | 621 | 36 | 644 |
| CAX68030.1 | 1.87e-258 | 11 | 628 | 31 | 655 |
| QGR35928.1 | 3.11e-232 | 12 | 621 | 15 | 620 |
| QPR29436.1 | 1.19e-222 | 13 | 630 | 17 | 627 |
| CBJ04457.1 | 1.24e-222 | 15 | 621 | 19 | 621 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3Q3E_A | 9.02e-120 | 20 | 614 | 23 | 613 | Crystalstructure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3E_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3H_A Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in complex with UDP-GLC [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3H_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in complex with UDP-GLC [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3I_A Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in the presence of peptide N1131 [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3I_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in the presence of peptide N1131 [Actinobacillus pleuropneumoniae serovar 1 str. 4074] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3N2T3 | 3.61e-119 | 20 | 614 | 12 | 602 | UDP-glucose:protein N-beta-glucosyltransferase OS=Actinobacillus pleuropneumoniae serotype 5b (strain L20) OX=416269 GN=APL_1635 PE=1 SV=1 |
| B3H2N2 | 2.70e-111 | 20 | 614 | 12 | 602 | UDP-glucose:protein N-beta-glucosyltransferase OS=Actinobacillus pleuropneumoniae serotype 7 (strain AP76) OX=537457 GN=APP7_1697 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
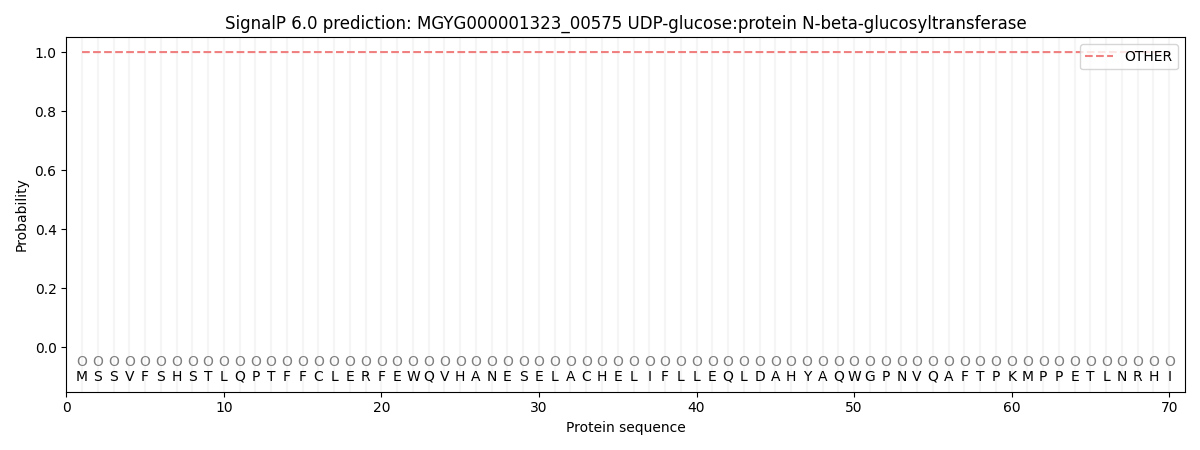
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000089 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
