You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001361_02285
You are here: Home > Sequence: MGYG000001361_02285
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sutterella wadsworthensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; Sutterella; Sutterella wadsworthensis | |||||||||||
| CAZyme ID | MGYG000001361_02285 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Cell division suppressor protein YneA | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 15404; End: 17515 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 114 | 245 | 1.3e-21 | 0.8444444444444444 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10783 | mltD | 1.01e-100 | 36 | 524 | 27 | 444 | membrane-bound lytic murein transglycosylase D; Provisional |
| cd16894 | MltD-like | 7.37e-59 | 118 | 246 | 1 | 128 | Membrane-bound lytic murein transglycosylase D and similar proteins. Lytic transglycosylases (LT) catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc). Membrane-bound lytic murein transglycosylase D protein (MltD) family members may have one or more small LysM domains, which may contribute to peptidoglycan binding. Unlike the similar "goose-type" lysozymes, LTs also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| PRK06347 | PRK06347 | 2.14e-29 | 470 | 702 | 318 | 591 | 1,4-beta-N-acetylmuramoylhydrolase. |
| PRK06347 | PRK06347 | 3.67e-28 | 338 | 581 | 328 | 591 | 1,4-beta-N-acetylmuramoylhydrolase. |
| pfam01464 | SLT | 1.01e-25 | 124 | 218 | 12 | 107 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQS89247.1 | 0.0 | 1 | 703 | 1 | 703 |
| QDA55156.1 | 0.0 | 45 | 703 | 37 | 702 |
| BBF22593.1 | 0.0 | 64 | 703 | 44 | 678 |
| ANU65188.1 | 2.93e-169 | 67 | 581 | 43 | 582 |
| QQQ96345.1 | 2.93e-169 | 67 | 581 | 43 | 582 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4B8V_A | 3.40e-13 | 526 | 703 | 21 | 217 | ChainA, Extracellular Protein 6 [Fulvia fulva],4B9H_A Chain A, Extracellular Protein 6 [Fulvia fulva] |
| 4UZ2_A | 8.49e-06 | 539 | 585 | 4 | 50 | Crystalstructure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_B Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_C Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ2_D Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4UZ3_A Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8],4UZ3_B Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8],4UZ3_C Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose [Thermus thermophilus HB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AEZ8 | 4.16e-64 | 77 | 462 | 70 | 446 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=mltD PE=3 SV=1 |
| P0AEZ7 | 4.16e-64 | 77 | 462 | 70 | 446 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli (strain K12) OX=83333 GN=mltD PE=1 SV=1 |
| P37710 | 9.16e-36 | 341 | 702 | 361 | 736 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| P32820 | 9.52e-29 | 106 | 218 | 21 | 134 | Putative tributyltin chloride resistance protein OS=Alteromonas sp. (strain M-1) OX=29457 GN=tbtA PE=3 SV=1 |
| O31852 | 4.66e-27 | 416 | 651 | 28 | 273 | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlS PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
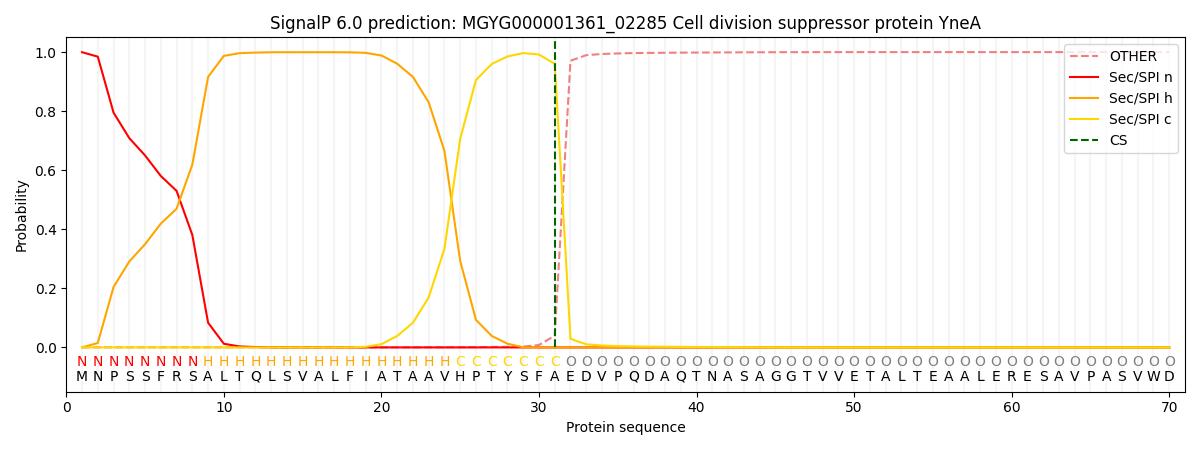
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001144 | 0.997895 | 0.000243 | 0.000311 | 0.000205 | 0.000167 |
