You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001574_00482
You are here: Home > Sequence: MGYG000001574_00482
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
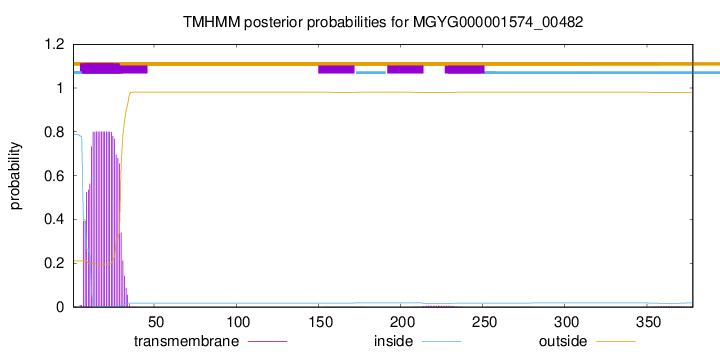
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; Sutterella; | |||||||||||
| CAZyme ID | MGYG000001574_00482 | |||||||||||
| CAZy Family | GH103 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase B | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 15638; End: 16774 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH103 | 99 | 374 | 5.2e-83 | 0.8983050847457628 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam13406 | SLT_2 | 2.70e-95 | 77 | 371 | 4 | 292 | Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. |
| TIGR02282 | MltB | 2.24e-85 | 62 | 373 | 1 | 290 | lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes. [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan] |
| COG2951 | MltB | 2.32e-81 | 1 | 378 | 1 | 340 | Membrane-bound lytic murein transglycosylase B [Cell wall/membrane/envelope biogenesis]. |
| PRK10760 | PRK10760 | 4.18e-66 | 50 | 376 | 49 | 356 | murein hydrolase B; Provisional |
| cd13399 | Slt35-like | 1.23e-17 | 154 | 272 | 1 | 74 | Slt35-like lytic transglycosylase. Lytic transglycosylase similar to Escherichia coli lytic transglycosylase Slt35 and Pseudomonas aeruginosa Sltb1. Lytic transglycosylase (LT) catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBF23912.1 | 7.13e-124 | 6 | 375 | 2 | 363 |
| QDA55336.1 | 1.88e-121 | 49 | 375 | 88 | 406 |
| QQS90106.1 | 4.90e-116 | 49 | 373 | 139 | 455 |
| ANU65602.1 | 1.11e-69 | 19 | 374 | 7 | 349 |
| QQQ96748.1 | 1.11e-69 | 19 | 374 | 7 | 349 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5O8X_A | 1.57e-52 | 118 | 376 | 51 | 302 | TheX-ray Structure of Catenated Lytic Transglycosylase SltB1 from Pseudomonas aeruginosa [Pseudomonas aeruginosa],5O8X_B The X-ray Structure of Catenated Lytic Transglycosylase SltB1 from Pseudomonas aeruginosa [Pseudomonas aeruginosa] |
| 4ANR_A | 2.47e-52 | 118 | 376 | 68 | 319 | Crystalstructure of soluble lytic Transglycosylase SltB1 from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1] |
| 1LTM_A | 7.25e-45 | 54 | 376 | 12 | 317 | AcceleratedX-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using Warp [Escherichia coli] |
| 1D0K_A | 7.62e-45 | 54 | 376 | 14 | 319 | ChainA, 35KD SOLUBLE LYTIC TRANSGLYCOSYLASE [Escherichia coli],1D0L_A Chain A, 35KD SOLUBLE LYTIC TRANSGLYCOSYLASE [Escherichia coli],1D0M_A Chain A, 35KD SOLUBLE LYTIC TRANSGLYCOSYLASE [Escherichia coli],1QDR_A Chain A, LYTIC MUREIN TRANSGLYCOSYLASE B [Escherichia coli],1QDT_A Chain A, LYTIC MUREIN TRANSGLYCOSYLASE B [Escherichia coli],1QUS_A Chain A, LYTIC MUREIN TRANSGLYCOSYLASE B [Escherichia coli],1QUT_A Chain A, LYTIC MUREIN TRANSGLYCOSYLASE B [Escherichia coli] |
| 5ANZ_A | 1.28e-28 | 122 | 371 | 98 | 341 | CrystalStructure of SltB3 from Pseudomonas aeruginosa. [Pseudomonas aeruginosa],5AO7_A Crystal Structure of SltB3 from Pseudomonas aeruginosa in complex with NAG-anhNAM-pentapeptide [Pseudomonas aeruginosa] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P41052 | 1.05e-43 | 54 | 376 | 53 | 358 | Membrane-bound lytic murein transglycosylase B OS=Escherichia coli (strain K12) OX=83333 GN=mltB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000355 | 0.998879 | 0.000175 | 0.000224 | 0.000184 | 0.000164 |