You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001657_00141
You are here: Home > Sequence: MGYG000001657_00141
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | RC9 sp900760425 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; RC9 sp900760425 | |||||||||||
| CAZyme ID | MGYG000001657_00141 | |||||||||||
| CAZy Family | GH125 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1246; End: 5100 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH125 | 871 | 1269 | 7.5e-178 | 0.9925373134328358 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3538 | COG3538 | 0.0 | 848 | 1271 | 2 | 422 | Meiotically up-regulated gene 157 (Mug157) protein (function unknown) [Function unknown]. |
| pfam06824 | Glyco_hydro_125 | 0.0 | 871 | 1269 | 3 | 416 | Metal-independent alpha-mannosidase (GH125). This family, which contains bacterial and fungal glycoside hydrolases, is also known as GH125. They function as metal-independent alpha-mannosidases, with specificity for alpha-1,6-linked non-reducing terminal mannose residues. Structurally this family is part of the 6 hairpin glycosidase superfamily. |
| pfam16335 | DUF4965 | 1.75e-93 | 486 | 655 | 1 | 174 | Domain of unknown function (DUF4965). This family consists of uncharacterized proteins around 840 residues in length and is mainly found in various Bacteroides species. Several proteins in this family are annotated as Glutaminases, but the function of this protein is unknown. |
| pfam08760 | DUF1793 | 2.46e-71 | 662 | 826 | 2 | 169 | Domain of unknown function (DUF1793). This presumed domain is found at the C-terminus of a glutaminase protein from fungi. This domain is also found as a single domain protein in Bacteroides thetaiotaomicron. |
| pfam17168 | DUF5127 | 9.82e-60 | 253 | 482 | 1 | 226 | Domain of unknown function (DUF5127). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCA48091.1 | 4.83e-259 | 140 | 826 | 11 | 697 |
| QUT73989.1 | 1.82e-221 | 843 | 1281 | 46 | 484 |
| QRO24558.1 | 4.94e-221 | 842 | 1281 | 42 | 481 |
| AUI48770.1 | 1.49e-220 | 843 | 1281 | 47 | 485 |
| QCQ35487.1 | 1.49e-220 | 843 | 1281 | 47 | 485 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2P0V_A | 1.66e-204 | 839 | 1281 | 39 | 478 | Crystalstructure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482],2P0V_B Crystal structure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482] |
| 3ON6_A | 3.39e-204 | 839 | 1281 | 19 | 458 | Crystalstructure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483],3ON6_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483] |
| 3P2C_A | 2.03e-203 | 843 | 1281 | 21 | 460 | Crystalstructure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483],3P2C_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483] |
| 6RQK_A | 5.99e-124 | 862 | 1278 | 18 | 429 | Crystalstructure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13],6RQK_B Crystal structure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13] |
| 5M7I_A | 3.15e-123 | 862 | 1278 | 18 | 429 | Crystalstructure of GH125 1,6-alpha-mannosidase mutant from Clostridium perfringens in complex with 1,6-alpha-mannobiose [Clostridium perfringens str. 13],5M7Y_A Crystal structure of GH125 1,6-alpha-mannosidase mutant from Clostridium perfringens in complex with 1,6-alpha-mannotriose [Clostridium perfringens str. 13] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q10449 | 1.75e-103 | 843 | 1282 | 51 | 506 | Meiotically up-regulated gene 157 protein OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mug157 PE=1 SV=1 |
| Q2U4L7 | 5.20e-73 | 245 | 826 | 97 | 689 | Glutaminase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=gtaA PE=1 SV=2 |
| D4AMT2 | 7.64e-41 | 185 | 822 | 56 | 779 | Probable glutaminase ARB_05535/05536 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_05535/05536 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
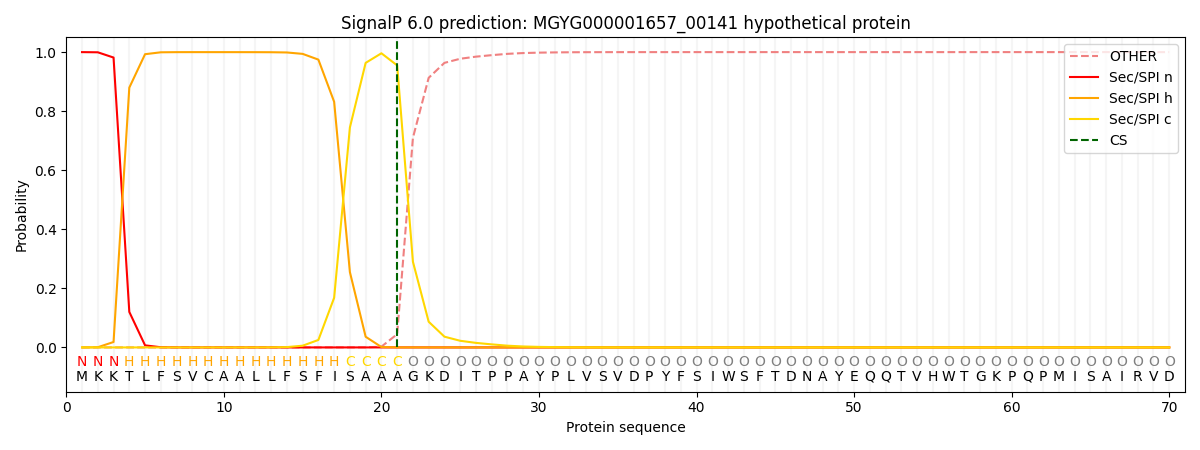
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000464 | 0.998642 | 0.000306 | 0.000212 | 0.000178 | 0.000164 |