You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001663_00769
You are here: Home > Sequence: MGYG000001663_00769
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp900760675 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp900760675 | |||||||||||
| CAZyme ID | MGYG000001663_00769 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 81871; End: 82560 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 64 | 223 | 9.7e-22 | 0.837037037037037 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13403 | MLTF-like | 3.29e-62 | 63 | 225 | 6 | 161 | membrane-bound lytic murein transglycosylase F (MLTF) and similar proteins. This subfamily includes membrane-bound lytic murein transglycosylase F (MltF, murein lyase F) that degrades murein glycan strands. It is responsible for catalyzing the release of 1,6-anhydromuropeptides from peptidoglycan. Lytic transglycosylase catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do goose-type lysozymes. However, in addition, it also makes a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| COG4623 | MltF | 3.03e-26 | 31 | 193 | 218 | 404 | Membrane-bound lytic murein transglycosylase MltF [Cell wall/membrane/envelope biogenesis, Signal transduction mechanisms]. |
| PRK10859 | PRK10859 | 8.98e-24 | 24 | 195 | 246 | 425 | membrane-bound lytic murein transglycosylase MltF. |
| pfam01464 | SLT | 4.67e-19 | 58 | 168 | 1 | 107 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd16896 | LT_Slt70-like | 5.15e-17 | 54 | 161 | 4 | 110 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFL78639.1 | 3.01e-137 | 1 | 229 | 1 | 230 |
| BBL08197.1 | 2.54e-134 | 1 | 229 | 1 | 232 |
| BBL10988.1 | 2.54e-134 | 1 | 229 | 1 | 232 |
| BBL00117.1 | 2.54e-134 | 1 | 229 | 1 | 232 |
| CBK65133.1 | 6.47e-132 | 1 | 229 | 1 | 230 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5AA4_B | 1.34e-17 | 54 | 221 | 252 | 412 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013],5AA4_D Crystal structure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013] |
| 5AA4_A | 1.34e-17 | 54 | 221 | 252 | 412 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013],5AA4_C Crystal structure of MltF from Pseudomonas aeruginosa in complex with cell-wall tetrapeptide [Pseudomonas aeruginosa BWHPSA013] |
| 5AA2_B | 1.48e-17 | 54 | 221 | 300 | 460 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with NAM-pentapeptide. [Pseudomonas aeruginosa BWHPSA013] |
| 5AA2_A | 1.48e-17 | 54 | 221 | 300 | 460 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with NAM-pentapeptide. [Pseudomonas aeruginosa BWHPSA013],5AA2_C Crystal structure of MltF from Pseudomonas aeruginosa in complex with NAM-pentapeptide. [Pseudomonas aeruginosa BWHPSA013] |
| 5AA2_D | 1.48e-17 | 54 | 221 | 300 | 460 | Crystalstructure of MltF from Pseudomonas aeruginosa in complex with NAM-pentapeptide. [Pseudomonas aeruginosa BWHPSA013] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0VRG6 | 5.51e-19 | 54 | 221 | 288 | 448 | Membrane-bound lytic murein transglycosylase F OS=Alcanivorax borkumensis (strain ATCC 700651 / DSM 11573 / NCIMB 13689 / SK2) OX=393595 GN=mltF PE=3 SV=1 |
| B4RVK5 | 3.48e-18 | 68 | 221 | 306 | 452 | Membrane-bound lytic murein transglycosylase F OS=Alteromonas mediterranea (strain DSM 17117 / CIP 110805 / LMG 28347 / Deep ecotype) OX=1774373 GN=mltF PE=3 SV=3 |
| Q4KHS7 | 3.70e-18 | 54 | 193 | 289 | 424 | Membrane-bound lytic murein transglycosylase F OS=Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5) OX=220664 GN=mltF PE=3 SV=2 |
| A1U3J1 | 7.05e-18 | 54 | 226 | 286 | 451 | Membrane-bound lytic murein transglycosylase F OS=Marinobacter nauticus (strain ATCC 700491 / DSM 11845 / VT8) OX=351348 GN=mltF PE=3 SV=1 |
| A5UG36 | 9.22e-18 | 48 | 193 | 271 | 422 | Membrane-bound lytic murein transglycosylase F OS=Haemophilus influenzae (strain PittGG) OX=374931 GN=mltF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
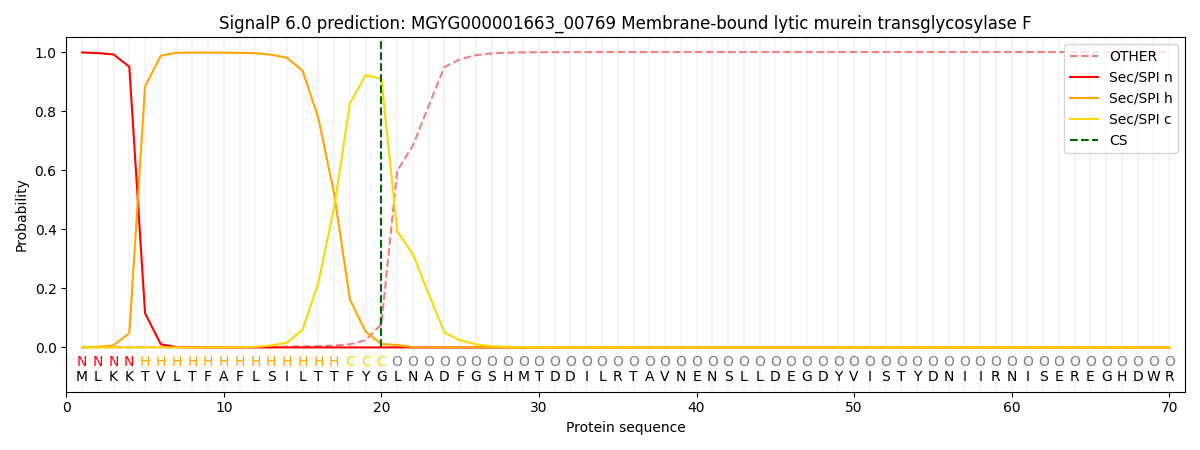
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.002789 | 0.995607 | 0.000884 | 0.000222 | 0.000227 | 0.000234 |
