You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001703_01429
You are here: Home > Sequence: MGYG000001703_01429
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Vibrio fluvialis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Vibrionaceae; Vibrio; Vibrio fluvialis | |||||||||||
| CAZyme ID | MGYG000001703_01429 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | Chitinase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 65371; End: 67920 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 159 | 578 | 2.4e-73 | 0.9459459459459459 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3325 | ChiA | 1.05e-170 | 123 | 592 | 2 | 441 | Chitinase, GH18 family [Carbohydrate transport and metabolism]. |
| cd06548 | GH18_chitinase | 1.13e-125 | 161 | 575 | 1 | 322 | The GH18 (glycosyl hydrolases, family 18) type II chitinases hydrolyze chitin, an abundant polymer of N-acetylglucosamine and have been identified in bacteria, fungi, insects, plants, viruses, and protozoan parasites. The structure of this domain is an eight-stranded alpha/beta barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. |
| smart00636 | Glyco_18 | 8.71e-125 | 160 | 575 | 1 | 334 | Glyco_18 domain. |
| pfam00704 | Glyco_hydro_18 | 1.87e-89 | 160 | 575 | 1 | 307 | Glycosyl hydrolases family 18. |
| pfam08329 | ChitinaseA_N | 1.43e-59 | 21 | 155 | 1 | 130 | Chitinase A, N-terminal domain. This domain is found in a number of bacterial chitinases and similar viral proteins. It is organized into a fibronectin III module domain-like fold, comprising only beta strands. Its function is not known, but it may be involved in interaction with the enzyme substrate, chitin. It is separated by a hinge region from the catalytic domain (pfam00704); this hinge region is probably mobile, allowing the N-terminal domain to have different relative positions in solution. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AMF93260.1 | 0.0 | 1 | 849 | 1 | 849 |
| QKE36713.1 | 0.0 | 1 | 849 | 1 | 849 |
| QUF70760.1 | 0.0 | 1 | 849 | 1 | 849 |
| AVH33468.1 | 0.0 | 1 | 849 | 1 | 846 |
| QTH10583.1 | 0.0 | 1 | 798 | 1 | 799 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3B9A_A | 0.0 | 22 | 590 | 1 | 569 | ChainA, Chitinase A [Vibrio harveyi],3B9D_A Chain A, Chitinase A [Vibrio harveyi],3B9E_A Chain A, Chitinase A [Vibrio harveyi] |
| 3ARO_A | 0.0 | 22 | 590 | 1 | 569 | CrystalStructure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - apo structure [Vibrio harveyi],3ARP_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with DEQUALINIUM [Vibrio harveyi],3ARQ_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with IDARUBICIN [Vibrio harveyi],3ARR_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with PENTOXIFYLLINE [Vibrio harveyi],3ARV_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with Sanguinarine [Vibrio harveyi],3ARW_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with chelerythrine [Vibrio harveyi],3ARX_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with Propentofylline [Vibrio harveyi],3ARY_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with 2-(imidazolin-2-yl)-5-isothiocyanatobenzofuran [Vibrio harveyi],3ARZ_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with 2-(imidazolin-2-yl)-5-isothiocyanatobenzofuran [Vibrio harveyi],3B8S_A Crystal structure of wild-type chitinase A from Vibrio harveyi [Vibrio harveyi],3B8S_B Crystal structure of wild-type chitinase A from Vibrio harveyi [Vibrio harveyi] |
| 3ARS_A | 0.0 | 22 | 590 | 1 | 569 | CrystalStructure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - apo structure of mutant W275G [Vibrio harveyi],3ART_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with DEQUALINIUM [Vibrio harveyi],3ARU_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with PENTOXIFYLLINE [Vibrio harveyi],3AS0_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with Sanguinarine [Vibrio harveyi],3AS1_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with chelerythrine [Vibrio harveyi],3AS2_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with Propentofylline [Vibrio harveyi],3AS3_A Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with 2-(imidazolin-2-yl)-5-isothiocyanatobenzofuran [Vibrio harveyi] |
| 2WK2_A | 8.09e-188 | 21 | 587 | 1 | 533 | ChitinaseA from Serratia marcescens ATCC990 in complex with Chitotrio-thiazoline dithioamide. [Serratia marcescens] |
| 2WLY_A | 1.06e-187 | 21 | 587 | 1 | 533 | ChitinaseA from Serratia marcescens ATCC990 in complex with Chitotrio-thiazoline. [Serratia marcescens],2WLZ_A Chitinase A from Serratia marcescens ATCC990 in complex with Chitobio- thiazoline. [Serratia marcescens],2WM0_A Chitinase A from Serratia marcescens ATCC990 in complex with Chitobio- thiazoline thioamide. [Serratia marcescens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P32823 | 1.96e-268 | 15 | 673 | 15 | 666 | Chitinase A OS=Pseudoalteromonas piscicida OX=43662 GN=chiA PE=1 SV=1 |
| P07254 | 1.55e-185 | 16 | 587 | 19 | 556 | Chitinase A OS=Serratia marcescens OX=615 GN=chiA PE=1 SV=3 |
| P41684 | 2.18e-151 | 21 | 584 | 17 | 542 | Chitinase OS=Autographa californica nuclear polyhedrosis virus OX=46015 GN=CHIA PE=1 SV=1 |
| O10363 | 8.08e-148 | 21 | 585 | 16 | 542 | Probable endochitinase OS=Orgyia pseudotsugata multicapsid polyhedrosis virus OX=262177 GN=ORF124 PE=3 SV=1 |
| P20533 | 1.17e-64 | 156 | 627 | 42 | 493 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
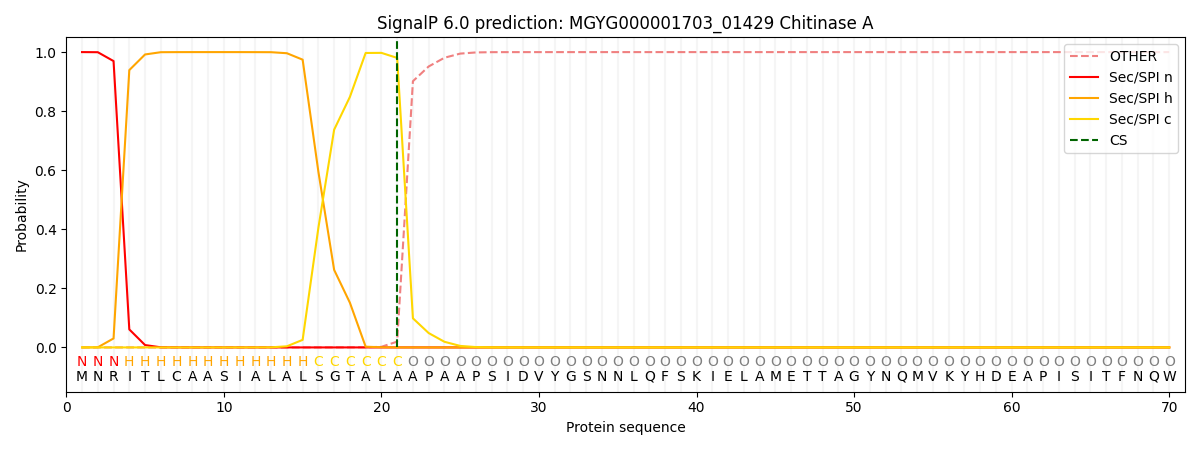
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000304 | 0.998942 | 0.000188 | 0.000203 | 0.000187 | 0.000168 |
