You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001775_00909
You are here: Home > Sequence: MGYG000001775_00909
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; Paramuribaculum; | |||||||||||
| CAZyme ID | MGYG000001775_00909 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 32002; End: 33039 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 62 | 297 | 7.1e-71 | 0.8062283737024222 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3934 | COG3934 | 1.59e-22 | 29 | 292 | 49 | 329 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| pfam09537 | DUF2383 | 8.19e-04 | 158 | 210 | 12 | 66 | Domain of unknown function (DUF2383). Members of this protein family are found mostly in the Proteobacteria, although one member is found in the the marine planctomycete Pirellula sp. strain 1. The function is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD38974.1 | 8.89e-113 | 19 | 301 | 36 | 317 |
| QCP72664.1 | 8.89e-113 | 19 | 301 | 36 | 317 |
| AGY54507.1 | 3.23e-105 | 29 | 301 | 31 | 303 |
| QJR73285.1 | 4.72e-105 | 20 | 293 | 26 | 298 |
| QJR68955.1 | 4.72e-105 | 20 | 293 | 26 | 298 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1UUQ_A | 9.12e-56 | 12 | 307 | 9 | 326 | Exo-mannosidasefrom Cellvibrio mixtus [Cellvibrio mixtus],1UZ4_A Common inhibition of beta-glucosidase and beta-mannosidase by isofagomine lactam reflects different conformational intineraries for glucoside and mannoside hydrolysis [Cellvibrio mixtus],7ODJ_AAA Chain AAA, Man5A [Cellvibrio mixtus] |
| 4LYR_A | 2.76e-51 | 19 | 286 | 34 | 298 | GlycosideHydrolase Family 5 Mannosidase from Rhizomucor miehei, E301A mutant [Rhizomucor miehei] |
| 4LYP_A | 2.76e-51 | 19 | 286 | 34 | 298 | CrystalStructure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei [Rhizomucor miehei],4LYP_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei [Rhizomucor miehei] |
| 4LYQ_A | 2.07e-50 | 19 | 286 | 34 | 298 | CrystalStructure of Glycoside Hydrolase Family 5 Mannosidase from Rhizomucor miehei, E202A mutant [Rhizomucor miehei],4NRR_A Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannosyl-fructose [Rhizomucor miehei],4NRR_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannosyl-fructose [Rhizomucor miehei],4NRS_A Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannobiose [Rhizomucor miehei],4NRS_B Crystal Structure of Glycoside Hydrolase Family 5 Mannosidase (E202A mutant) from Rhizomucor miehei in complex with mannobiose [Rhizomucor miehei] |
| 3PZ9_A | 1.44e-27 | 39 | 312 | 27 | 308 | Nativestructure of endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3PZG_A I222 crystal form of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3PZI_A Structure of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 in complex with beta-D-glucose [Thermotoga petrophila RKU-1],3PZM_A Structure of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 with three glycerol molecules [Thermotoga petrophila RKU-1],3PZN_A Structure of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 with citrate and glycerol [Thermotoga petrophila RKU-1],3PZO_A Structure of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 in complex with three maltose molecules [Thermotoga petrophila RKU-1],3PZQ_A Structure of the hyperthermostable endo-1,4-beta-D-mannanase from Thermotoga petrophila RKU-1 with maltose and glycerol [Thermotoga petrophila RKU-1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q10B67 | 1.81e-29 | 26 | 295 | 24 | 288 | Mannan endo-1,4-beta-mannosidase 4 OS=Oryza sativa subsp. japonica OX=39947 GN=MAN4 PE=2 SV=2 |
| Q9SG95 | 8.44e-28 | 63 | 259 | 66 | 246 | Putative mannan endo-1,4-beta-mannosidase 4 OS=Arabidopsis thaliana OX=3702 GN=MAN4 PE=3 SV=1 |
| Q9FZ29 | 8.77e-28 | 43 | 285 | 48 | 281 | Mannan endo-1,4-beta-mannosidase 1 OS=Arabidopsis thaliana OX=3702 GN=MAN1 PE=2 SV=1 |
| Q5W6G0 | 1.07e-27 | 43 | 300 | 97 | 359 | Putative mannan endo-1,4-beta-mannosidase 5 OS=Oryza sativa subsp. japonica OX=39947 GN=MAN5 PE=2 SV=2 |
| Q6Z310 | 1.15e-27 | 43 | 285 | 58 | 288 | Putative mannan endo-1,4-beta-mannosidase 9 OS=Oryza sativa subsp. japonica OX=39947 GN=MAN9 PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
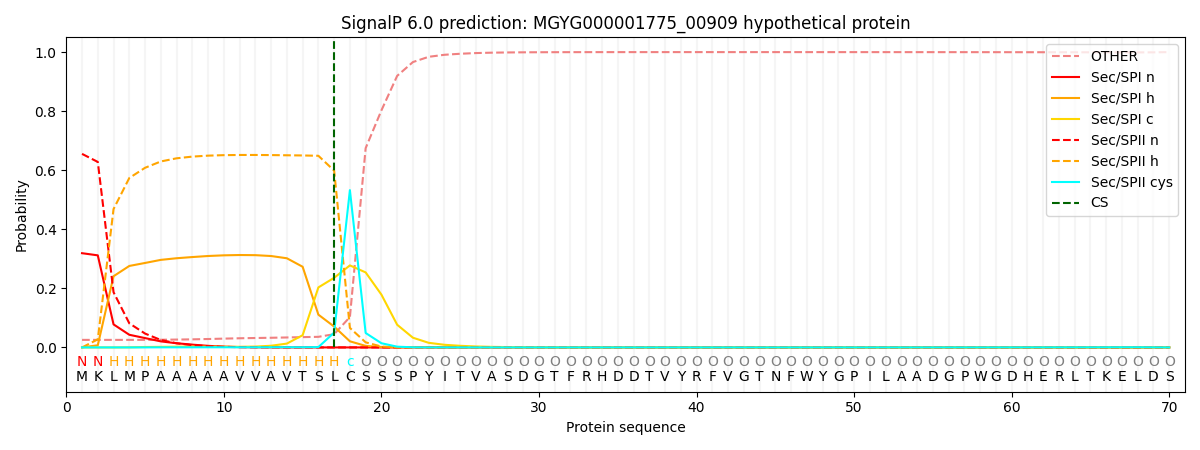
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.028818 | 0.311315 | 0.659276 | 0.000195 | 0.000189 | 0.000205 |
