You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001921_01037
You are here: Home > Sequence: MGYG000001921_01037
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Verrucomicrobiales; Akkermansiaceae; Akkermansia; | |||||||||||
| CAZyme ID | MGYG000001921_01037 | |||||||||||
| CAZy Family | GT10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 93437; End: 94435 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT10 | 109 | 263 | 1e-39 | 0.4322766570605187 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00852 | Glyco_transf_10 | 1.13e-12 | 152 | 257 | 19 | 129 | Glycosyltransferase family 10 (fucosyltransferase) C-term. This is the C-terminal domain of a family of fucosyltransferases. This enzyme transfers fucose from GDP-Fucose to GlcNAc in an alpha1,3 linkage. This family is known as glycosyltransferase family 10. The C-terminal domain is the likely binding-region for ADP (manuscript in publication). |
| pfam18025 | FucT_N | 0.002 | 14 | 101 | 11 | 88 | Alpha-(1,3)-fucosyltransferase FucT N-terminal domain. This is the N-terminal domain of the alpha chain found in Helicobacter pylori Fucosyltransferase protein which is involved in the production of Lewis x trisaccharide, a major component of lipopolysaccharide. The N-terminal domain contains the catalyst base, Glu-95 which is equivalent to the Asp-100 of other members of the glycosyltransferases-B family. The domain contains the pocket where LacNAc binds. The domain is composed of 2-10 heptad repeats and a conserved N-terminal alpha-beta-alpha motif which has little sequence similarity to the conserved N-terminal motif in other glycosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNB42929.1 | 7.68e-255 | 1 | 332 | 1 | 332 |
| QUF78618.1 | 4.78e-249 | 1 | 332 | 1 | 332 |
| QWO89162.1 | 4.78e-249 | 1 | 332 | 1 | 332 |
| AYR32183.1 | 4.78e-249 | 1 | 332 | 1 | 332 |
| AYR31431.1 | 4.78e-249 | 1 | 332 | 1 | 332 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2NZW_A | 3.60e-23 | 72 | 275 | 95 | 322 | CrystalStructure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_C Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZX_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZY_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori] |
| 5ZOI_A | 5.18e-22 | 72 | 275 | 95 | 322 | CrystalStructure of alpha1,3-Fucosyltransferase [Helicobacter pylori],5ZOI_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5UQ63 | 1.17e-22 | 81 | 265 | 382 | 566 | Putative fucosyltransferase R654 OS=Acanthamoeba polyphaga mimivirus OX=212035 GN=MIMI_R654 PE=3 SV=1 |
| O30511 | 4.79e-22 | 72 | 275 | 95 | 322 | Alpha-(1,3)-fucosyltransferase FucT OS=Helicobacter pylori OX=210 GN=fucT PE=1 SV=1 |
| Q9C8W3 | 1.84e-08 | 73 | 260 | 139 | 359 | Alpha-(1,4)-fucosyltransferase OS=Arabidopsis thaliana OX=3702 GN=FUT13 PE=2 SV=2 |
| Q6NTZ6 | 2.08e-07 | 141 | 267 | 216 | 344 | Alpha-(1,3)-fucosyltransferase 10 OS=Xenopus laevis OX=8355 GN=fut10 PE=2 SV=2 |
| Q6A1G3 | 4.94e-07 | 141 | 267 | 218 | 346 | Alpha-(1,3)-fucosyltransferase 10 OS=Xenopus tropicalis OX=8364 GN=fut10 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
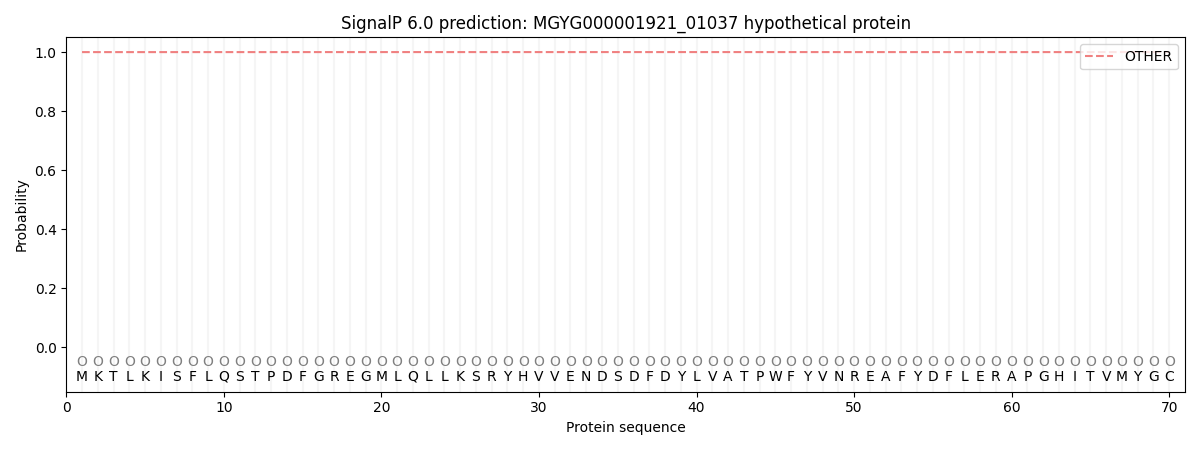
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000044 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
