You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001931_01845
You are here: Home > Sequence: MGYG000001931_01845
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
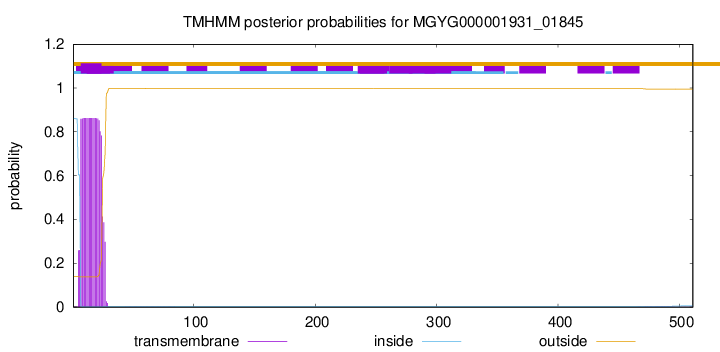
TMHMM annotations
Basic Information help
| Species | CAG-279 sp900548875 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-279; CAG-279 sp900548875 | |||||||||||
| CAZyme ID | MGYG000001931_01845 | |||||||||||
| CAZy Family | GH128 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1164; End: 2699 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam11790 | Glyco_hydro_cc | 2.24e-56 | 48 | 274 | 17 | 235 | Glycosyl hydrolase catalytic core. This family is probably a glycosyl hydrolase, and is conserved in fungi and some Proteobacteria. The pombe member is annotated as being from IPR013781. |
| TIGR04183 | Por_Secre_tail | 5.10e-07 | 453 | 510 | 14 | 71 | Por secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber average twenty or more copies of a C-terminal domain, represented by this model, associated with sorting to the outer membrane and covalent modification. |
| pfam18962 | Por_Secre_tail | 0.001 | 447 | 510 | 6 | 72 | Secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber have on average twenty or more copies of this C-terminal domain, associated with sorting to the outer membrane and covalent modification. This domain targets proteins to type IX secretion systems and is secreted then cleaved off by a C-terminal signal peptidease. Based on similarity to other families it is likely that this domain adopts an immunoglobulin like fold. |
| cd04079 | CBM6_agarase-like | 0.003 | 319 | 411 | 36 | 119 | Carbohydrate Binding Module 6 (CBM6); appended mainly to glycoside hydrolase (GH) family 16 alpha- and beta agarases. This family includes carbohydrate binding module 6 (CBM6) domains that are appended mainly to glycoside hydrolase (GH) family 16 agarases. These CBM6s are non-catalytic carbohydrate binding domains that facilitate the activity of alpha- and beta-agarase catalytic modules which are involved in the hydrolysis of 1,4-beta-D-galactosidic linkages. These CBM6s bind specifically to the non-reducing end of agarose chains, recognizing only the first repeat of the disaccharide, and directing the appended catalytic modules to areas of the plant cell wall attacked by beta-agarases. CBM6 is an unusual CBM as it represents a chimera of two distinct binding sites with different modes of binding: binding site I within the loop regions and binding site II on the concave face of the beta-sandwich fold. This family includes three tandem CBM6s from the Saccharophagus degradans agarase Aga86E, and three tandem CBM6s from Vibrio sp. strain PO-303 AgaA; in both these proteins these are appended to a GH16 domain. Vibrio AgaA also contains a Big-2-like protein-protein interaction domain. This family also includes two tandem CBM6s from an endo-type beta-agarase from a deep-sea Microbulbifer-like isolate, which are appended to a GH16 domain, and two of three CBM6s of Alteromonas agarilytica AgaA alpha-agarase, which are appended to a GH96 domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIU93826.1 | 2.39e-68 | 25 | 408 | 51 | 428 |
| ACR13446.1 | 1.47e-54 | 25 | 347 | 724 | 1041 |
| QEY18783.1 | 3.07e-53 | 20 | 407 | 731 | 1101 |
| QKX17146.1 | 1.52e-51 | 23 | 353 | 738 | 1063 |
| ABD82184.1 | 1.54e-51 | 24 | 407 | 734 | 1102 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6UAW_A | 7.67e-48 | 15 | 275 | 14 | 276 | Crystalstructure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Pseudomonas viridiflava (PvGH128_II) in complex with laminaritriose [Pseudomonas viridiflava] |
| 6UAV_A | 2.15e-47 | 25 | 275 | 2 | 254 | Crystalstructure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Pseudomonas viridiflava (PvGH128_II) [Pseudomonas viridiflava] |
| 6UAX_A | 4.40e-37 | 28 | 275 | 95 | 343 | Crystalstructure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Sorangium cellulosum (ScGH128_II) [Sorangium cellulosum So ce56],6UAX_B Crystal structure of a GH128 (subgroup II) endo-beta-1,3-glucanase from Sorangium cellulosum (ScGH128_II) [Sorangium cellulosum So ce56] |
| 6UAQ_A | 1.42e-21 | 48 | 260 | 44 | 254 | Crystalstructure of a GH128 (subgroup I) endo-beta-1,3-glucanase from Amycolatopsis mediterranei (AmGH128_I) [Amycolatopsis mediterranei],6UAR_A Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase from Amycolatopsis mediterranei (AmGH128_I) in complex with laminaritriose [Amycolatopsis mediterranei] |
| 6UFL_A | 3.56e-21 | 48 | 260 | 44 | 254 | Crystalstructure of a GH128 (subgroup I) endo-beta-1,3-glucanase (E199Q mutant) from Amycolatopsis mediterranei (AmGH128_I) in the complex with laminarihexaose [Amycolatopsis mediterranei],6UFZ_A Crystal structure of a GH128 (subgroup I) endo-beta-1,3-glucanase (E199Q mutant) from Amycolatopsis mediterranei (AmGH128_I) [Amycolatopsis mediterranei] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
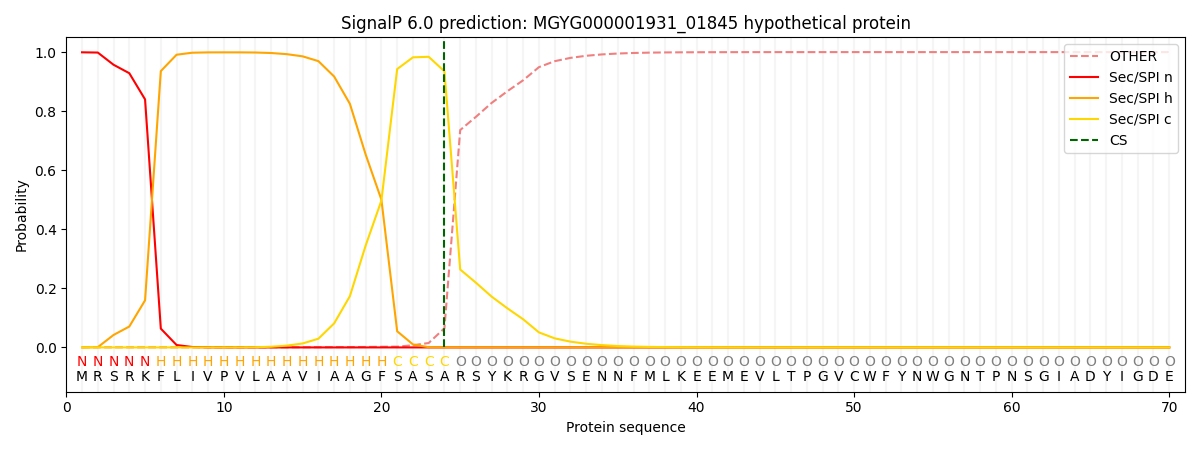
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001420 | 0.997157 | 0.000669 | 0.000315 | 0.000217 | 0.000189 |