You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002282_00719
You are here: Home > Sequence: MGYG000002282_00719
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides gordonii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides gordonii | |||||||||||
| CAZyme ID | MGYG000002282_00719 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | Xylan 1,3-beta-xylosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 872182; End: 873720 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 27 | 302 | 2.1e-122 | 0.99644128113879 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd08999 | GH43_ABN-like | 1.58e-89 | 27 | 303 | 1 | 284 | Glycosyl hydrolase family 43 protein such as endo-alpha-L-arabinanase. This glycosyl hydrolase family 43 (GH43) subgroup includes mostly enzymes with alpha-L-arabinofuranosidase (ABF; EC 3.2.1.55) and endo-alpha-L-arabinanase (ABN; EC 3.2.1.99) activities. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. The GH43 ABN enzymes hydrolyze alpha-1,5-L-arabinofuranoside linkages while the ABF enzymes cleave arabinose side chains so that the combined actions of these two enzymes reduce arabinan to L-arabinose and/or arabinooligosaccharides. These arabinan-degrading enzymes are important in the food industry for efficient production of L-arabinose from agricultural waste; L-arabinose is often used as a bioactive sweetener. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| pfam04616 | Glyco_hydro_43 | 3.13e-84 | 25 | 302 | 1 | 281 | Glycosyl hydrolases family 43. The glycosyl hydrolase family 43 contains members that are arabinanases. Arabinanases hydrolyze the alpha-1,5-linked L-arabinofuranoside backbone of plant cell wall arabinans. The structure of arabinanase Arb43A from Cellvibrio japonicus reveals a five-bladed beta-propeller fold. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd08991 | GH43_HoAraf43-like | 6.64e-63 | 35 | 300 | 1 | 276 | Glycosyl hydrolase family 43 protein such as Halothermothrix orenii H 168 alpha-L-arabinofuranosidase (HoAraf43;Hore_20580). This glycosyl hydrolase family 43 (GH43) subgroup includes Halothermothrix orenii H 168 alpha-L-arabinofuranosidase (EC 3.2.1.55) (HoAraf43;Hore_20580). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. This GH43_ HoAraf43-like subgroup includes enzymes that have been annotated as having xylan-digesting beta-xylosidase (EC 3.2.1.37) and xylanase (endo-alpha-L-arabinanase, EC 3.2.1.8) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18616 | GH43_ABN-like | 3.11e-57 | 27 | 286 | 1 | 277 | Glycosyl hydrolase family 43 such as arabinan endo-1 5-alpha-L-arabinosidase. This glycosyl hydrolase family 43 (GH43) subgroup includes mostly enzymes with endo-alpha-L-arabinanase (ABN; EC 3.2.1.99) activity. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. The GH43 ABN enzymes hydrolyze alpha-1,5-L-arabinofuranoside linkages. These arabinan-degrading enzymes are important in the food industry for efficient production of L-arabinose from agricultural waste; L-arabinose is often used as a bioactive sweetener. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG3507 | XynB2 | 1.67e-55 | 1 | 496 | 1 | 531 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUR47469.1 | 4.80e-306 | 1 | 512 | 1 | 513 |
| QUT55603.1 | 6.81e-306 | 1 | 512 | 1 | 513 |
| QCY57382.1 | 9.67e-306 | 1 | 512 | 1 | 513 |
| QIX64229.1 | 1.95e-305 | 1 | 512 | 1 | 513 |
| QJE30638.1 | 2.77e-305 | 1 | 512 | 1 | 513 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MS3_A | 6.19e-32 | 17 | 466 | 23 | 485 | Crystalstructure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580],6MS3_B Crystal structure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580] |
| 6MS2_A | 1.14e-31 | 17 | 466 | 23 | 485 | Crystalstructure of the GH43 BlXynB protein from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580] |
| 2EXH_A | 1.42e-29 | 27 | 480 | 6 | 502 | Structureof the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_B Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_C Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_D Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 2EXJ_A | 1.92e-29 | 27 | 480 | 6 | 502 | ChainA, beta-D-xylosidase [Geobacillus stearothermophilus],2EXJ_B Chain B, beta-D-xylosidase [Geobacillus stearothermophilus],2EXJ_C Chain C, beta-D-xylosidase [Geobacillus stearothermophilus],2EXJ_D Chain D, beta-D-xylosidase [Geobacillus stearothermophilus] |
| 2EXK_A | 1.17e-28 | 27 | 480 | 6 | 502 | ChainA, beta-D-xylosidase [Geobacillus stearothermophilus],2EXK_B Chain B, beta-D-xylosidase [Geobacillus stearothermophilus],2EXK_C Chain C, beta-D-xylosidase [Geobacillus stearothermophilus],2EXK_D Chain D, beta-D-xylosidase [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94489 | 3.81e-27 | 26 | 339 | 4 | 339 | Beta-xylosidase OS=Bacillus subtilis (strain 168) OX=224308 GN=xynB PE=1 SV=2 |
| A9ZND1 | 1.28e-26 | 23 | 478 | 3 | 495 | Xylan 1,3-beta-xylosidase OS=Vibrio sp. OX=678 GN=xloA PE=1 SV=1 |
| P95470 | 1.71e-26 | 33 | 302 | 35 | 331 | Extracellular exo-alpha-(1->5)-L-arabinofuranosidase ArbA OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=arbA PE=1 SV=1 |
| A7LXT8 | 2.36e-25 | 25 | 366 | 23 | 363 | Non-reducing end alpha-L-arabinofuranosidase BoGH43A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02654 PE=1 SV=1 |
| Q93HT9 | 2.57e-22 | 36 | 303 | 27 | 312 | Intracellular endo-alpha-(1->5)-L-arabinanase OS=Geobacillus thermodenitrificans OX=33940 GN=abn-ts PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
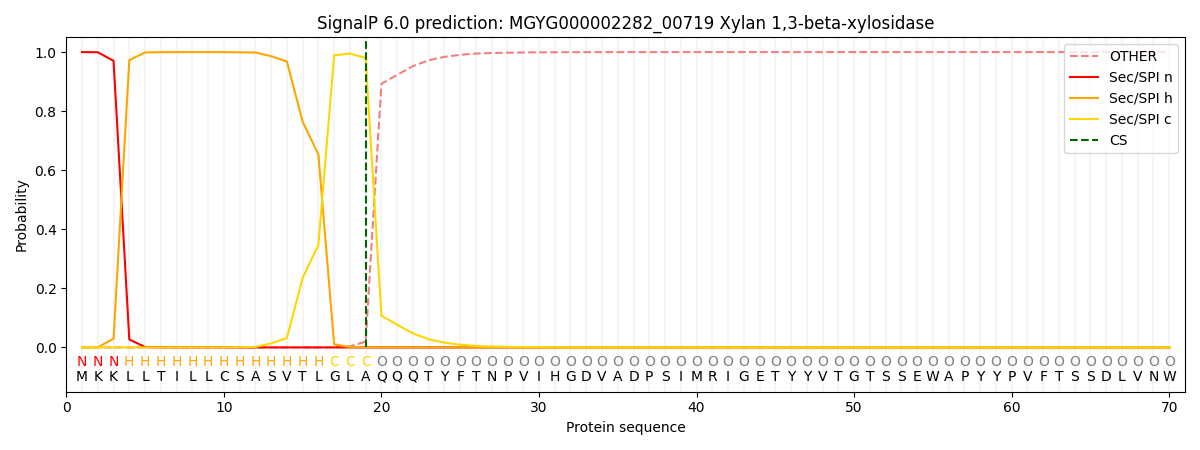
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000282 | 0.999030 | 0.000189 | 0.000171 | 0.000158 | 0.000143 |
