You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002398_01660
You are here: Home > Sequence: MGYG000002398_01660
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Laribacter hongkongensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Aquaspirillaceae; Laribacter; Laribacter hongkongensis | |||||||||||
| CAZyme ID | MGYG000002398_01660 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Soluble lytic murein transglycosylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1698064; End: 1699980 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13401 | Slt70-like | 8.06e-61 | 459 | 603 | 1 | 150 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| cd16896 | LT_Slt70-like | 1.18e-49 | 463 | 598 | 3 | 143 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| PRK11619 | PRK11619 | 5.99e-49 | 21 | 602 | 38 | 624 | lytic murein transglycosylase; Provisional |
| COG0741 | MltE | 3.32e-33 | 323 | 606 | 3 | 287 | Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Cell wall/membrane/envelope biogenesis]. |
| cd00254 | LT-like | 2.97e-32 | 479 | 599 | 1 | 110 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ASJ24577.1 | 0.0 | 1 | 638 | 1 | 638 |
| ACO74476.1 | 0.0 | 1 | 638 | 1 | 638 |
| AVY95345.1 | 1.06e-241 | 9 | 638 | 15 | 645 |
| AQR64946.1 | 6.38e-206 | 13 | 622 | 12 | 617 |
| BAK77109.1 | 9.97e-186 | 13 | 622 | 16 | 624 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5O2O_A | 7.09e-126 | 6 | 622 | 10 | 615 | Lytictransglycosylase in action [Neisseria meningitidis],6H5F_A LtgA disordered Helix [Neisseria meningitidis NM422] |
| 5MPQ_A | 5.31e-124 | 40 | 622 | 8 | 575 | BulgecinA: The key to a broad-spectrum inhibitor that targets lytic transglycosylases [Neisseria meningitidis] |
| 5O1J_A | 5.98e-124 | 40 | 622 | 12 | 579 | Lytictransglycosylase in action [Neisseria meningitidis MC58] |
| 6FPN_B | 7.14e-124 | 40 | 622 | 18 | 585 | Lytictransglycosylase in action [Neisseria meningitidis MC58] |
| 5O24_A | 8.04e-124 | 40 | 622 | 22 | 589 | Lytictransglycosylase in action [Neisseria meningitidis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39434 | 1.45e-42 | 31 | 602 | 47 | 625 | Soluble lytic murein transglycosylase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=slt PE=3 SV=2 |
| P0AGC3 | 1.43e-40 | 31 | 615 | 47 | 638 | Soluble lytic murein transglycosylase OS=Escherichia coli (strain K12) OX=83333 GN=slt PE=1 SV=1 |
| P0AGC4 | 1.43e-40 | 31 | 615 | 47 | 638 | Soluble lytic murein transglycosylase OS=Escherichia coli O157:H7 OX=83334 GN=slt PE=3 SV=1 |
| P44888 | 1.46e-17 | 298 | 602 | 273 | 572 | Putative soluble lytic murein transglycosylase OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=slt PE=3 SV=1 |
| O31976 | 2.70e-13 | 460 | 598 | 1418 | 1540 | SPbeta prophage-derived uncharacterized transglycosylase YomI OS=Bacillus subtilis (strain 168) OX=224308 GN=yomI PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
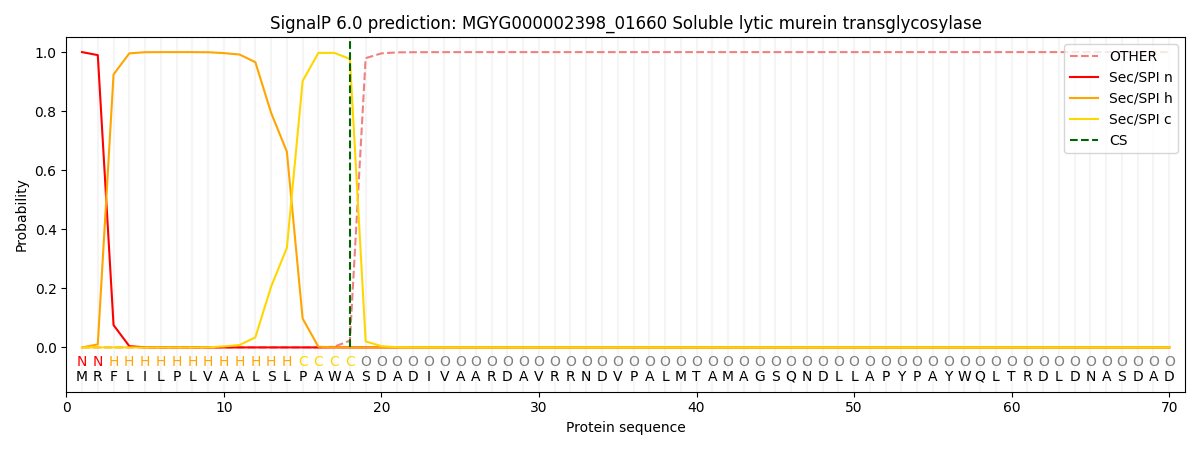
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000398 | 0.998835 | 0.000215 | 0.000181 | 0.000161 | 0.000162 |
