You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002454_00856
You are here: Home > Sequence: MGYG000002454_00856
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Akkermansia muciniphila | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Verrucomicrobiales; Akkermansiaceae; Akkermansia; Akkermansia muciniphila | |||||||||||
| CAZyme ID | MGYG000002454_00856 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | Choline-sulfatase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1035141; End: 1036826 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16144 | ARS_like | 9.78e-179 | 36 | 522 | 1 | 418 | uncharacterized arylsulfatase subfamily. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16145 | ARS_like | 3.79e-92 | 36 | 512 | 1 | 415 | uncharacterized arylsulfatase subfamily. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16026 | GALNS_like | 5.63e-87 | 35 | 508 | 1 | 399 | galactosamine-6-sulfatase; also known as N-acetylgalactosamine-6-sulfatase (GALNS). Lysosomal galactosamine-6-sulfatase removes sulfate groups from a terminal N-acetylgalactosamine-6-sulfate (or galactose-6-sulfate) in mucopolysaccharides such as keratan sulfate and chondroitin-6-sulfate. Defects in GALNS lead to accumulation of substrates, resulting in the development of the lysosomal storage disease mucopolysaccharidosis IV A. |
| cd16146 | ARS_like | 8.68e-84 | 36 | 518 | 1 | 397 | uncharacterized arylsulfatase. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16029 | 4-S | 2.10e-75 | 36 | 508 | 1 | 393 | N-acetylgalactosamine 4-sulfatase, also called arylsulftase B. Sulfatases catalyze the hydrolysis of sulfuric acid esters from a wide variety of substrates. N-acetylgalactosamine 4-sulfatase catalyzes the removal of the sulfate ester group from position 4 of an N-acetylgalactosamine sugar at the non-reducing terminus of the polysaccharide in the degradative pathways of the glycosaminoglycans dermatan sulfate and chondroitin-4-sulfate. N-acetylgalactosamine 4-sulfatase is a lysosomal enzyme. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| EAR02039.1 | 1.45e-131 | 35 | 556 | 28 | 511 |
| QUT89376.1 | 6.40e-130 | 17 | 554 | 6 | 498 |
| ALJ59588.1 | 9.88e-129 | 29 | 554 | 17 | 498 |
| AKJ65363.1 | 7.17e-58 | 34 | 551 | 25 | 467 |
| QEG00402.1 | 2.01e-41 | 7 | 518 | 712 | 1154 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6USS_A | 5.05e-133 | 35 | 556 | 33 | 517 | ChainA, Sulfatase [Bacteroides fragilis CAG:558],6USS_B Chain B, Sulfatase [Bacteroides fragilis CAG:558] |
| 6UST_A | 1.11e-69 | 35 | 534 | 4 | 461 | ChainA, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_B Chain B, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_C Chain C, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_D Chain D, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi] |
| 7STT_A | 6.68e-58 | 34 | 534 | 5 | 441 | ChainA, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis],7STU_A Chain A, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis],7STV_A Chain A, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis] |
| 6BIA_A | 2.64e-50 | 27 | 508 | 18 | 425 | Crystalstructure of Ps i-CgsB [Pseudoalteromonas fuliginea],6BIA_B Crystal structure of Ps i-CgsB [Pseudoalteromonas fuliginea],6BIA_C Crystal structure of Ps i-CgsB [Pseudoalteromonas fuliginea] |
| 6B0K_A | 7.59e-48 | 34 | 508 | 1 | 401 | Crystalstructure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas],6B0K_B Crystal structure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas],6B0K_C Crystal structure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q32KH5 | 1.59e-38 | 15 | 456 | 12 | 386 | N-acetylgalactosamine-6-sulfatase OS=Canis lupus familiaris OX=9615 GN=GALNS PE=2 SV=1 |
| T2KPJ9 | 1.74e-38 | 34 | 557 | 33 | 549 | Sulfatase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22020 PE=3 SV=1 |
| Q571E4 | 8.96e-37 | 34 | 458 | 26 | 386 | N-acetylgalactosamine-6-sulfatase OS=Mus musculus OX=10090 GN=Galns PE=1 SV=2 |
| P34059 | 1.71e-36 | 34 | 456 | 29 | 386 | N-acetylgalactosamine-6-sulfatase OS=Homo sapiens OX=9606 GN=GALNS PE=1 SV=1 |
| Q32KJ6 | 2.40e-36 | 34 | 458 | 30 | 390 | N-acetylgalactosamine-6-sulfatase OS=Rattus norvegicus OX=10116 GN=Galns PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
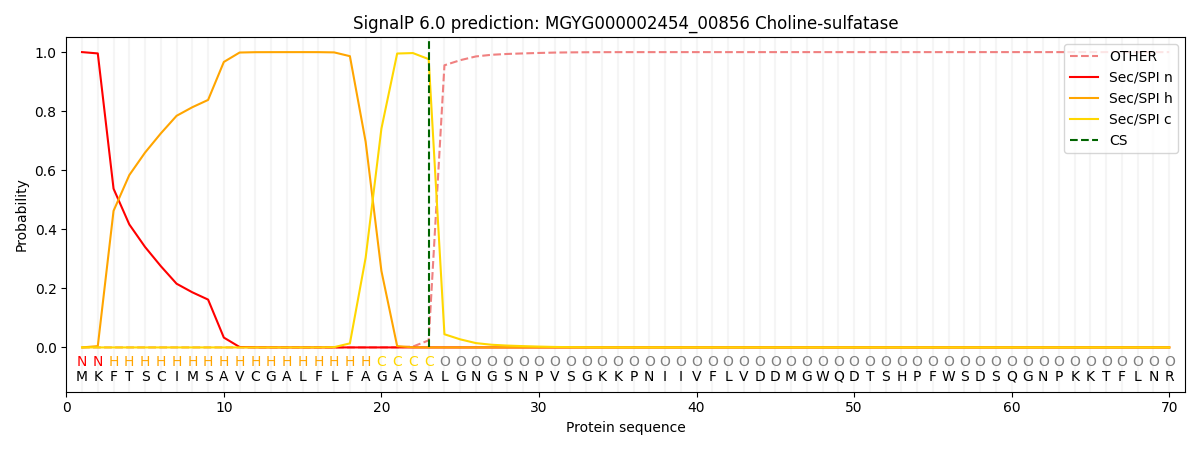
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000403 | 0.998857 | 0.000172 | 0.000208 | 0.000169 | 0.000149 |
