You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002460_03972
You are here: Home > Sequence: MGYG000002460_03972
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
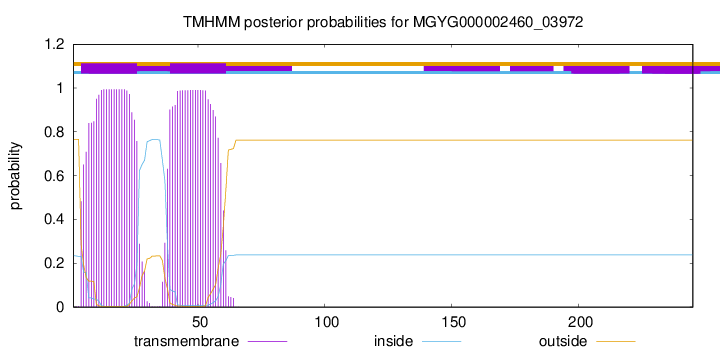
TMHMM annotations
Basic Information help
| Species | Cronobacter sakazakii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Cronobacter; Cronobacter sakazakii | |||||||||||
| CAZyme ID | MGYG000002460_03972 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4185560; End: 4186297 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 93 | 229 | 5.9e-18 | 0.7481481481481481 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13403 | MLTF-like | 1.15e-27 | 95 | 222 | 16 | 137 | membrane-bound lytic murein transglycosylase F (MLTF) and similar proteins. This subfamily includes membrane-bound lytic murein transglycosylase F (MltF, murein lyase F) that degrades murein glycan strands. It is responsible for catalyzing the release of 1,6-anhydromuropeptides from peptidoglycan. Lytic transglycosylase catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do goose-type lysozymes. However, in addition, it also makes a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd13399 | Slt35-like | 9.18e-22 | 90 | 182 | 4 | 96 | Slt35-like lytic transglycosylase. Lytic transglycosylase similar to Escherichia coli lytic transglycosylase Slt35 and Pseudomonas aeruginosa Sltb1. Lytic transglycosylase (LT) catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| cd16896 | LT_Slt70-like | 6.13e-17 | 71 | 229 | 2 | 142 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| COG4623 | MltF | 8.28e-17 | 83 | 232 | 282 | 436 | Membrane-bound lytic murein transglycosylase MltF [Cell wall/membrane/envelope biogenesis, Signal transduction mechanisms]. |
| cd00254 | LT-like | 3.13e-16 | 94 | 228 | 4 | 107 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AXX04240.1 | 3.73e-191 | 1 | 245 | 1 | 245 |
| AST79516.1 | 3.73e-191 | 1 | 245 | 1 | 245 |
| BAE74105.1 | 3.73e-191 | 1 | 245 | 1 | 245 |
| ASV20922.1 | 2.16e-190 | 1 | 245 | 1 | 245 |
| QHF78700.1 | 2.16e-190 | 1 | 245 | 1 | 245 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1SLY_A | 3.03e-08 | 30 | 185 | 416 | 560 | ComplexOf The 70-Kda Soluble Lytic Transglycosylase With Bulgecin A [Escherichia coli] |
| 1QSA_A | 2.38e-07 | 30 | 185 | 416 | 560 | CrystalStructure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.65 Angstroms Resolution [Escherichia coli],1QTE_A Crystal Structure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.90 A Resolution In Complex With A 1,6- Anhydromurotripeptide [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A1U3J1 | 2.24e-10 | 76 | 202 | 292 | 406 | Membrane-bound lytic murein transglycosylase F OS=Marinobacter nauticus (strain ATCC 700491 / DSM 11845 / VT8) OX=351348 GN=mltF PE=3 SV=1 |
| Q2SK06 | 5.51e-10 | 83 | 202 | 315 | 422 | Membrane-bound lytic murein transglycosylase F OS=Hahella chejuensis (strain KCTC 2396) OX=349521 GN=mltF PE=3 SV=1 |
| A1S850 | 1.77e-09 | 94 | 201 | 299 | 400 | Membrane-bound lytic murein transglycosylase F OS=Shewanella amazonensis (strain ATCC BAA-1098 / SB2B) OX=326297 GN=mltF PE=3 SV=2 |
| B0TLJ6 | 3.18e-09 | 94 | 201 | 296 | 397 | Membrane-bound lytic murein transglycosylase F OS=Shewanella halifaxensis (strain HAW-EB4) OX=458817 GN=mltF PE=3 SV=1 |
| A8H256 | 5.79e-09 | 94 | 201 | 296 | 397 | Membrane-bound lytic murein transglycosylase F OS=Shewanella pealeana (strain ATCC 700345 / ANG-SQ1) OX=398579 GN=mltF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
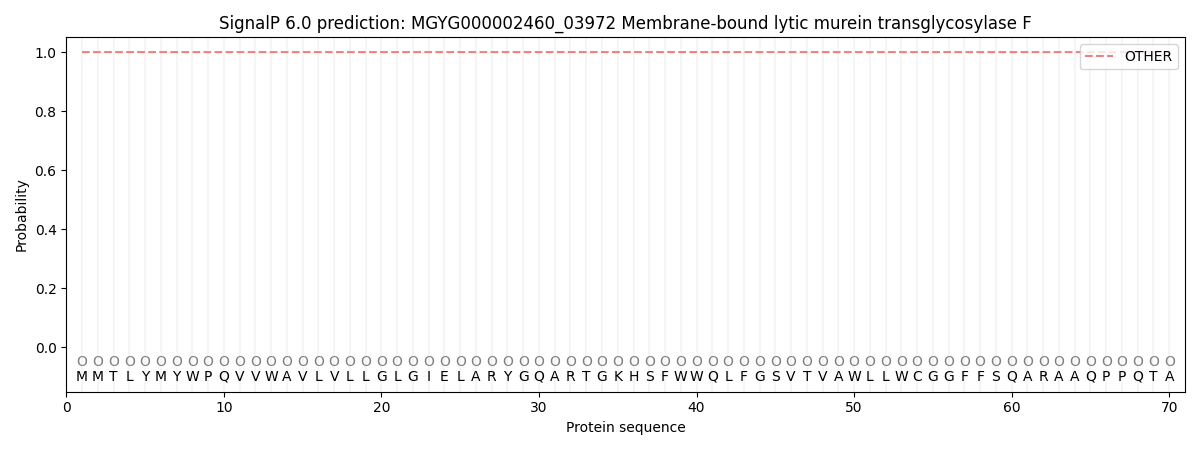
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000017 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |