You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002490_04993
You are here: Home > Sequence: MGYG000002490_04993
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_B thiaminolyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_B; Paenibacillus_B thiaminolyticus | |||||||||||
| CAZyme ID | MGYG000002490_04993 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | Endo-beta-N-acetylglucosaminidase F2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 226771; End: 227751 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06542 | GH18_EndoS-like | 2.23e-57 | 44 | 311 | 1 | 253 | Endo-beta-N-acetylglucosaminidases are bacterial chitinases that hydrolyze the chitin core of various asparagine (N)-linked glycans and glycoproteins. The endo-beta-N-acetylglucosaminidases have a glycosyl hydrolase family 18 (GH18) catalytic domain. Some members also have an additional C-terminal glycosyl hydrolase family 20 (GH20) domain while others have an N-terminal domain of unknown function (pfam08522). Members of this family include endo-beta-N-acetylglucosaminidase S (EndoS) from Streptococcus pyogenes, EndoF1, EndoF2, EndoF3, and EndoH from Flavobacterium meningosepticum, and EndoE from Enterococcus faecalis. EndoS is a secreted endoglycosidase from Streptococcus pyogenes that specifically hydrolyzes the glycan on human IgG between two core N-acetylglucosamine residues. EndoE is a secreted endoglycosidase, encoded by the ndoE gene in Enterococcus faecalis, that hydrolyzes the glycan on human RNase B. |
| pfam16141 | DUF4849 | 7.46e-08 | 63 | 299 | 45 | 301 | Putative glycoside hydrolase Family 18, chitinase_18. This DUF is likely to be a form of glycosyl hydrolase from CAZy family 18, possibly chitinase 18. This would have the EC number of EC:3.2.1.14. |
| cd00598 | GH18_chitinase-like | 6.98e-07 | 46 | 226 | 1 | 176 | The GH18 (glycosyl hydrolase, family 18) type II chitinases hydrolyze chitin, an abundant polymer of beta-1,4-linked N-acetylglucosamine (GlcNAc) which is a major component of the cell wall of fungi and the exoskeleton of arthropods. Chitinases have been identified in viruses, bacteria, fungi, protozoan parasites, insects, and plants. The structure of the GH18 domain is an eight-stranded beta/alpha barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. The GH18 family includes chitotriosidase, chitobiase, hevamine, zymocin-alpha, narbonin, SI-CLP (stabilin-1 interacting chitinase-like protein), IDGF (imaginal disc growth factor), CFLE (cortical fragment-lytic enzyme) spore hydrolase, the type III and type V plant chitinases, the endo-beta-N-acetylglucosaminidases, and the chitolectins. The GH85 (glycosyl hydrolase, family 85) ENGases (endo-beta-N-acetylglucosaminidases) are closely related to the GH18 chitinases and are included in this alignment model. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDM44162.1 | 2.26e-244 | 1 | 326 | 1 | 326 |
| SYX83265.1 | 1.34e-213 | 3 | 325 | 2 | 324 |
| ALS02650.1 | 2.05e-153 | 1 | 325 | 1 | 317 |
| ALS36881.1 | 2.19e-151 | 46 | 323 | 1 | 279 |
| CCO12584.1 | 7.35e-151 | 1 | 324 | 1 | 315 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6KPL_A | 2.08e-73 | 32 | 323 | 16 | 295 | CrystalStructure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris in apo form [Cordyceps militaris CM01],6KPM_A Crystal Structure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris in complex with L-fucose [Cordyceps militaris CM01] |
| 6KPN_A | 3.32e-72 | 32 | 323 | 16 | 295 | CrystalStructure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris D154N/E156Q mutant in complex with fucosyl-N-acetylglucosamine [Cordyceps militaris CM01],6KPO_A Crystal Structure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris D154N/E156Q mutant in complex with fucosyl-N-acetylglucosamine-Asn [Cordyceps militaris CM01] |
| 7PUJ_A | 9.49e-64 | 35 | 320 | 5 | 291 | ChainA, Beta-N-acetylhexosaminidase [Enterococcus faecalis],7PUK_A Chain A, Beta-N-acetylhexosaminidase [Enterococcus faecalis],7PUK_C Chain C, Beta-N-acetylhexosaminidase [Enterococcus faecalis] |
| 6E58_A | 1.66e-43 | 34 | 319 | 13 | 338 | Crystalstructure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) [Streptococcus pyogenes M49 591],6E58_B Crystal structure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) [Streptococcus pyogenes M49 591],6MDS_A Crystal structure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) with complex biantennary glycan [Streptococcus pyogenes],6MDS_B Crystal structure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) with complex biantennary glycan [Streptococcus pyogenes],6MDV_A Crystal structure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) with high-mannose glycan [Streptococcus pyogenes],6MDV_B Crystal structure of Streptococcus pyogenes endo-beta-N-acetylglucosaminidase (EndoS2) with high-mannose glycan [Streptococcus pyogenes] |
| 4NUY_A | 3.82e-37 | 44 | 326 | 15 | 351 | Crystalstructure of EndoS, an endo-beta-N-acetyl-glucosaminidase from Streptococcus pyogenes [Streptococcus pyogenes serotype M1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P36912 | 1.36e-57 | 45 | 323 | 62 | 333 | Endo-beta-N-acetylglucosaminidase F2 OS=Elizabethkingia meningoseptica OX=238 GN=endOF2 PE=1 SV=1 |
| B7GPC7 | 5.10e-51 | 47 | 320 | 54 | 341 | Endo-beta-N-acetylglucosaminidase OS=Bifidobacterium longum subsp. infantis (strain ATCC 15697 / DSM 20088 / JCM 1222 / NCTC 11817 / S12) OX=391904 GN=Blon_2468 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
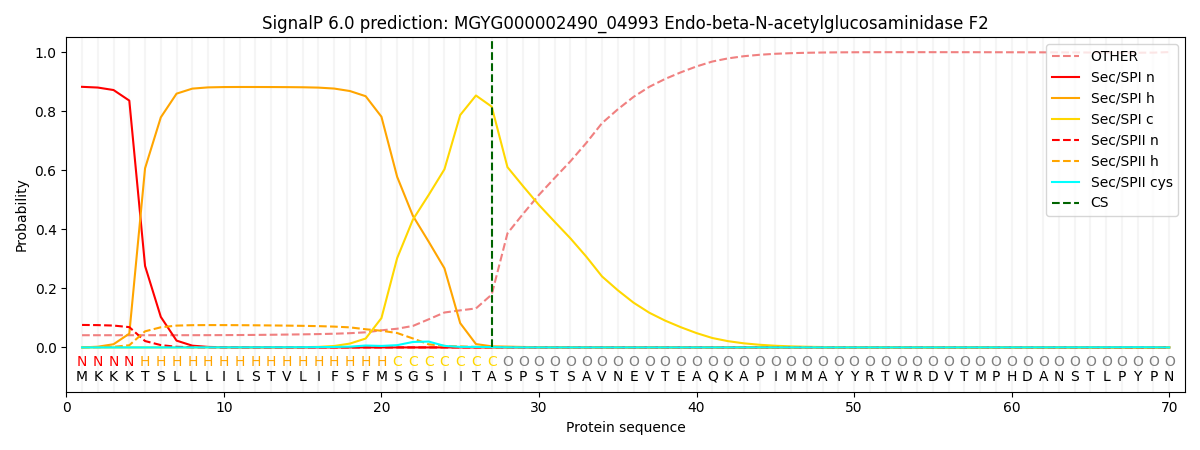
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.049309 | 0.869831 | 0.079316 | 0.000781 | 0.000367 | 0.000365 |

