You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002506_00358
You are here: Home > Sequence: MGYG000002506_00358
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Escherichia coli_D | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Escherichia; Escherichia coli_D | |||||||||||
| CAZyme ID | MGYG000002506_00358 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Glucans biosynthesis glucosyltransferase H | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 367028; End: 369541 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 236 | 418 | 1.6e-24 | 0.9764705882352941 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2943 | MdoH | 0.0 | 45 | 822 | 1 | 736 | Membrane glycosyltransferase [Cell wall/membrane/envelope biogenesis, Carbohydrate transport and metabolism]. |
| PRK05454 | PRK05454 | 0.0 | 62 | 706 | 1 | 599 | glucans biosynthesis glucosyltransferase MdoH. |
| cd04191 | Glucan_BSP_MdoH | 7.56e-155 | 234 | 487 | 1 | 254 | Glucan_BSP_MdoH catalyzes the elongation of beta-1,2 polyglucose chains of glucan. Periplasmic Glucan Biosynthesis protein MdoH is a glucosyltransferase that catalyzes the elongation of beta-1,2 polyglucose chains of glucan, requiring a beta-glucoside as a primer and UDP-glucose as a substrate. Glucans are composed of 5 to 10 units of glucose forming a highly branched structure, where beta-1,2-linked glucose constitutes a linear backbone to which branches are attached by beta-1,6 linkages. In Escherichia coli, glucans are located in the periplasmic space, functioning as regulator of osmolarity. It is synthesized at a maximum when cells are grown in a medium with low osmolarity. It has been shown to span the cytoplasmic membrane. |
| pfam00535 | Glycos_transf_2 | 7.49e-12 | 237 | 416 | 3 | 164 | Glycosyl transferase family 2. Diverse family, transferring sugar from UDP-glucose, UDP-N-acetyl- galactosamine, GDP-mannose or CDP-abequose, to a range of substrates including cellulose, dolichol phosphate and teichoic acids. |
| pfam13632 | Glyco_trans_2_3 | 6.80e-11 | 331 | 531 | 1 | 194 | Glycosyl transferase family group 2. Members of this family of prokaryotic proteins include putative glucosyltransferases, which are involved in bacterial capsule biosynthesis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIL17706.1 | 0.0 | 1 | 837 | 1 | 837 |
| QHW41423.1 | 0.0 | 1 | 837 | 11 | 847 |
| VWQ01965.1 | 0.0 | 1 | 837 | 1 | 837 |
| AZH89259.1 | 0.0 | 1 | 837 | 11 | 847 |
| AZH53718.1 | 0.0 | 1 | 837 | 11 | 847 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q48PK6 | 0.0 | 35 | 828 | 51 | 843 | Glucans biosynthesis glucosyltransferase H OS=Pseudomonas savastanoi pv. phaseolicola (strain 1448A / Race 6) OX=264730 GN=opgH PE=3 SV=1 |
| Q6D6A7 | 0.0 | 15 | 822 | 35 | 847 | Glucans biosynthesis glucosyltransferase H OS=Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672) OX=218491 GN=mdoH PE=3 SV=1 |
| Q4ZZH4 | 0.0 | 1 | 828 | 18 | 843 | Glucans biosynthesis glucosyltransferase H OS=Pseudomonas syringae pv. syringae (strain B728a) OX=205918 GN=opgH PE=3 SV=1 |
| Q4KJM5 | 0.0 | 19 | 837 | 37 | 854 | Glucans biosynthesis glucosyltransferase H OS=Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5) OX=220664 GN=opgH PE=3 SV=1 |
| Q82SA8 | 0.0 | 1 | 831 | 11 | 841 | Glucans biosynthesis glucosyltransferase H OS=Nitrosomonas europaea (strain ATCC 19718 / CIP 103999 / KCTC 2705 / NBRC 14298) OX=228410 GN=opgH PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
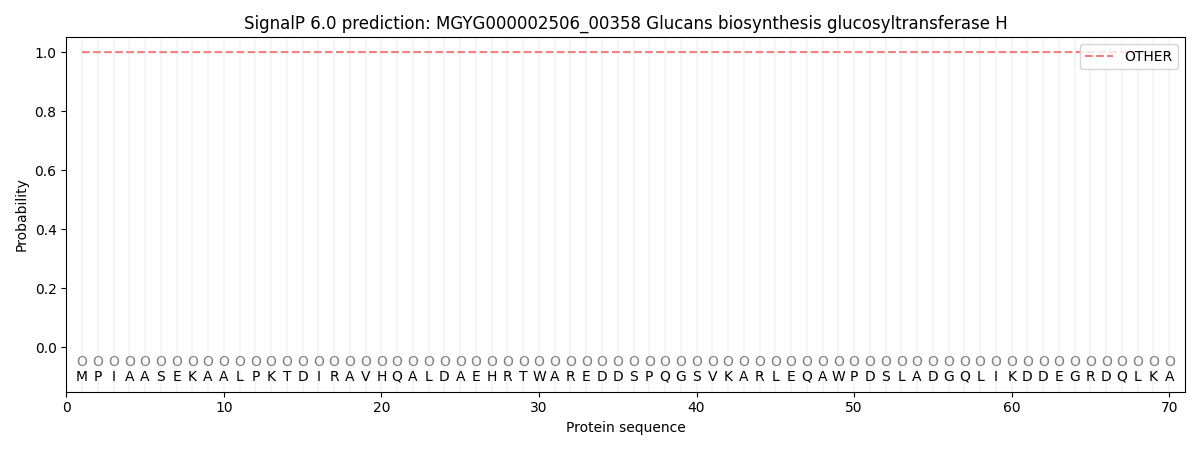
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999930 | 0.000121 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
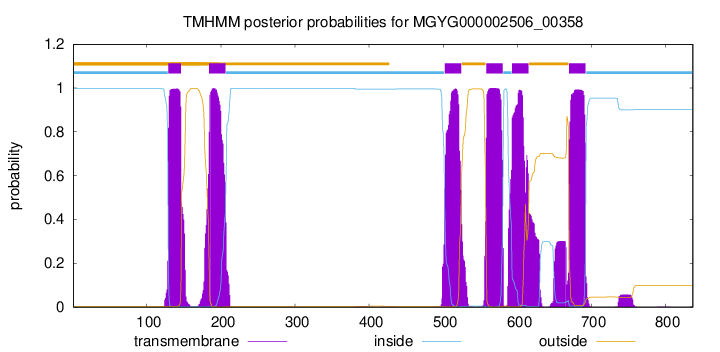
| start | end |
|---|---|
| 129 | 146 |
| 184 | 206 |
| 502 | 524 |
| 558 | 580 |
| 593 | 615 |
| 670 | 692 |
