You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002513_01853
You are here: Home > Sequence: MGYG000002513_01853
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Morganella morganii_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Morganella; Morganella morganii_A | |||||||||||
| CAZyme ID | MGYG000002513_01853 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | Glycosyltransferase Gtf1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 33520; End: 34596 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK09922 | PRK09922 | 2.00e-121 | 6 | 352 | 2 | 348 | lipopolysaccharide 1,6-galactosyltransferase. |
| cd03811 | GT4_GT28_WabH-like | 1.26e-60 | 16 | 349 | 10 | 349 | family 4 and family 28 glycosyltransferases similar to Klebsiella WabH. This family is most closely related to the GT1 family of glycosyltransferases. WabH in Klebsiella pneumoniae has been shown to transfer a GlcNAc residue from UDP-GlcNAc onto the acceptor GalUA residue in the cellular outer core. |
| cd03820 | GT4_AmsD-like | 1.88e-43 | 6 | 318 | 1 | 312 | amylovoran biosynthesis glycosyltransferase AmsD and similar proteins. This family is most closely related to the GT4 family of glycosyltransferases. AmSD in Erwinia amylovora has been shown to be involved in the biosynthesis of amylovoran, the acidic exopolysaccharide acting as a virulence factor. This enzyme may be responsible for the formation of galactose alpha-1,6 linkages in amylovoran. |
| cd03808 | GT4_CapM-like | 1.08e-30 | 82 | 333 | 72 | 334 | capsular polysaccharide biosynthesis glycosyltransferase CapM and similar proteins. This family is most closely related to the GT4 family of glycosyltransferases. CapM in Staphylococcus aureus is required for the synthesis of type 1 capsular polysaccharides. |
| COG0438 | RfaB | 5.48e-25 | 7 | 327 | 4 | 341 | Glycosyltransferase involved in cell wall bisynthesis [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QXO78386.1 | 3.59e-264 | 1 | 358 | 1 | 358 |
| QXO82073.1 | 3.59e-264 | 1 | 358 | 1 | 358 |
| QXO51610.1 | 3.59e-264 | 1 | 358 | 1 | 358 |
| QXO59419.1 | 3.59e-264 | 1 | 358 | 1 | 358 |
| QXO63160.1 | 3.59e-264 | 1 | 358 | 1 | 358 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5N7Z_A | 4.83e-86 | 6 | 349 | 2 | 345 | glycosyltransferasein LPS biosynthesis [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],6Y6G_A Chain A, Lipopolysaccharide 1,6-galactosyltransferase [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5N80_A | 4.98e-86 | 6 | 349 | 3 | 346 | glycosyltransferaseLPS biosynthesis in complex with UDP [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 6Y6I_A | 5.13e-86 | 6 | 349 | 4 | 347 | ChainA, Lipopolysaccharide 1,6-galactosyltransferase [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P27127 | 8.27e-87 | 6 | 356 | 2 | 350 | Lipopolysaccharide 1,6-galactosyltransferase OS=Escherichia coli (strain K12) OX=83333 GN=rfaB PE=3 SV=2 |
| Q06994 | 2.64e-85 | 6 | 349 | 2 | 345 | Lipopolysaccharide 1,6-galactosyltransferase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=rfaB PE=1 SV=2 |
| Q46634 | 5.92e-17 | 6 | 329 | 3 | 320 | Amylovoran biosynthesis glycosyltransferase AmsD OS=Erwinia amylovora OX=552 GN=amsD PE=3 SV=2 |
| O05083 | 1.66e-12 | 13 | 328 | 10 | 322 | Uncharacterized glycosyltransferase HI_1698 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=HI_1698 PE=3 SV=1 |
| Q58459 | 3.92e-11 | 20 | 306 | 11 | 321 | Uncharacterized glycosyltransferase MJ1059 OS=Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440) OX=243232 GN=MJ1059 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
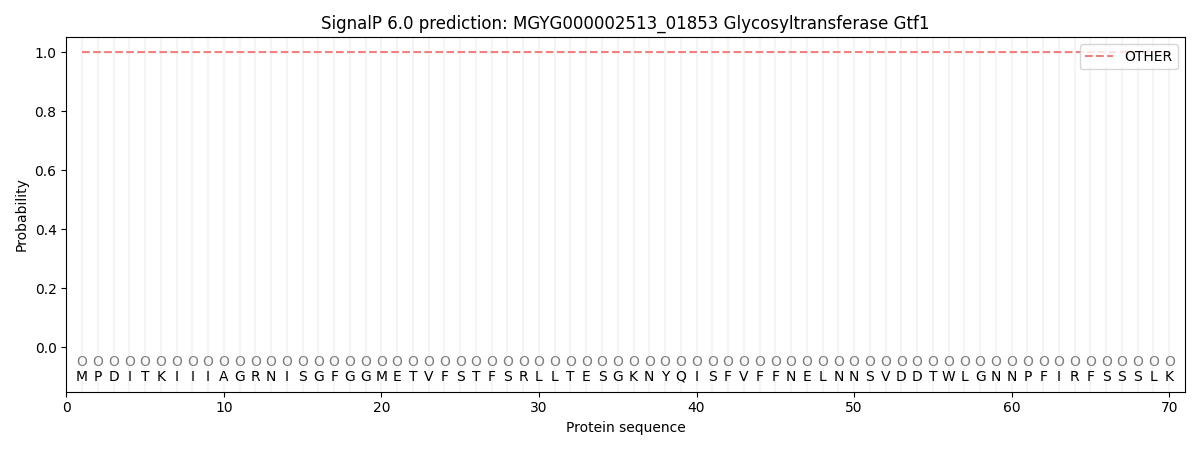
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000069 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
