You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002757_00572
You are here: Home > Sequence: MGYG000002757_00572
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Collinsella sp900550595 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; Collinsella sp900550595 | |||||||||||
| CAZyme ID | MGYG000002757_00572 | |||||||||||
| CAZy Family | GH105 | |||||||||||
| CAZyme Description | Unsaturated rhamnogalacturonyl hydrolase YteR | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 57199; End: 58437 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH105 | 33 | 374 | 2.5e-107 | 0.9728915662650602 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4225 | YesR | 2.23e-88 | 3 | 372 | 16 | 352 | Rhamnogalacturonyl hydrolase YesR [Carbohydrate transport and metabolism]. |
| pfam07470 | Glyco_hydro_88 | 1.51e-77 | 33 | 372 | 22 | 338 | Glycosyl Hydrolase Family 88. Unsaturated glucuronyl hydrolase catalyzes the hydrolytic release of unsaturated glucuronic acids from oligosaccharides (EC:3.2.1.-) produced by the reactions of polysaccharide lyases. |
| COG3533 | COG3533 | 1.56e-04 | 42 | 123 | 180 | 262 | Uncharacterized conserved protein, DUF1680 family [Function unknown]. |
| pfam07944 | Glyco_hydro_127 | 0.004 | 43 | 124 | 178 | 261 | Beta-L-arabinofuranosidase, GH127. One member of this family, from Bidobacterium longicum, UniProtKB:E8MGH8, has been characterized as an unusual beta-L-arabinofuranosidase enzyme, EC:3.2.1.185. It rleases l-arabinose from the l-arabinofuranose (Araf)-beta1,2-Araf disaccharide and also transglycosylates 1-alkanols with retention of the anomeric configuration. Terminal beta-l-arabinofuranosyl residues have been found in arabinogalactan proteins from a mumber of different plantt species. beta-l-Arabinofuranosyl linkages with 1-4 arabinofuranosides are also found in the sugar chains of extensin and solanaceous lectins, hydroxyproline (Hyp)2-rich glycoproteins that are widely observed in plant cell wall fractions. The critical residue for catalytic activity is Glu-338, in a ET/SCAS sequence context. |
| COG1331 | YyaL | 0.005 | 45 | 77 | 473 | 509 | Uncharacterized conserved protein YyaL, SSP411 family, contains thoiredoxin and six-hairpin glycosidase-like domains [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIA34435.1 | 0.0 | 1 | 412 | 1 | 412 |
| ATP54720.1 | 9.83e-317 | 1 | 412 | 1 | 412 |
| QOL34305.1 | 6.91e-229 | 1 | 383 | 1 | 381 |
| QOL31037.1 | 4.64e-227 | 1 | 383 | 1 | 381 |
| QOV20148.1 | 1.25e-193 | 5 | 374 | 2 | 377 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4WU0_A | 1.46e-91 | 38 | 372 | 23 | 357 | StructuralAnalysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824],4WU0_B Structural Analysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824] |
| 1NC5_A | 3.70e-58 | 36 | 382 | 37 | 373 | Structureof Protein of Unknown Function of YteR from Bacillus Subtilis [Bacillus subtilis],2D8L_A Crystal Structure of Unsaturated Rhamnogalacturonyl Hydrolase in complex with dGlcA-GalNAc [Bacillus subtilis] |
| 2GH4_A | 1.56e-57 | 36 | 382 | 27 | 363 | ChainA, Putative glycosyl hydrolase yteR [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34559 | 2.03e-57 | 36 | 382 | 37 | 373 | Unsaturated rhamnogalacturonyl hydrolase YteR OS=Bacillus subtilis (strain 168) OX=224308 GN=yteR PE=1 SV=1 |
| P0A3U6 | 3.31e-38 | 149 | 374 | 1 | 229 | Protein Atu3128 OS=Agrobacterium fabrum (strain C58 / ATCC 33970) OX=176299 GN=Atu3128 PE=3 SV=1 |
| P0A3U7 | 3.31e-38 | 149 | 374 | 1 | 229 | 24.9 kDa protein in picA locus OS=Rhizobium radiobacter OX=358 PE=2 SV=1 |
| O31521 | 7.21e-21 | 38 | 327 | 29 | 297 | Unsaturated rhamnogalacturonyl hydrolase YesR OS=Bacillus subtilis (strain 168) OX=224308 GN=yesR PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
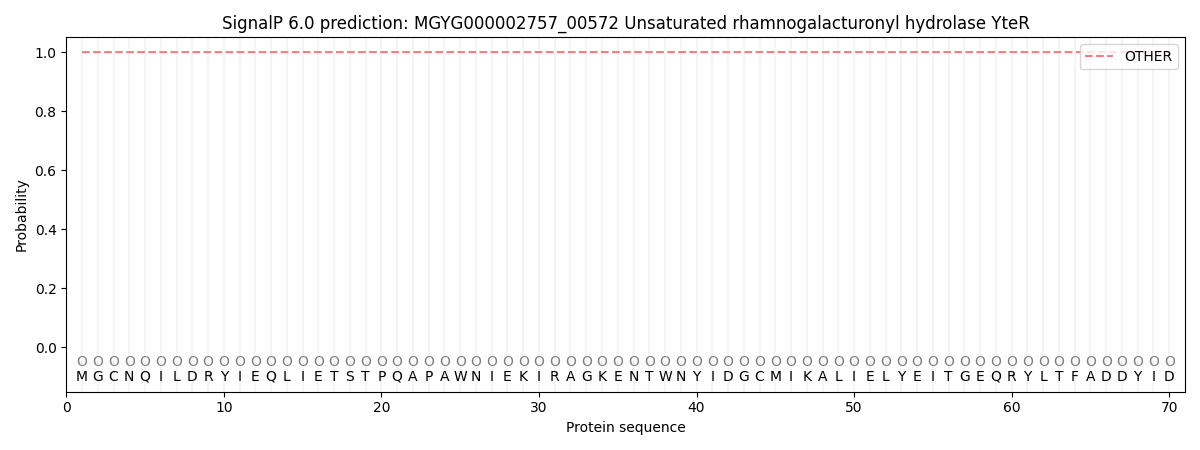
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
