You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002762_00548
You are here: Home > Sequence: MGYG000002762_00548
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-485 sp900554845 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; CAG-485 sp900554845 | |||||||||||
| CAZyme ID | MGYG000002762_00548 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | Xylosidase/arabinosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10793; End: 12394 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 26 | 338 | 2.1e-103 | 0.9931972789115646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd18620 | GH43_XylA-like | 8.46e-109 | 36 | 351 | 1 | 274 | Glycosyl hydrolase family 43-like protein such as Clostridium stercorarium alpha-L-arabinofuranosidase XylA. This glycosyl hydrolase family 43 (GH43) subgroup belongs to the GH43_AXH-like subgroup which includes enzymes that have been characterized with beta-xylosidase (EC 3.2.1.37), alpha-L-arabinofuranosidase (EC 3.2.1.55), alpha-1,2-L-arabinofuranosidase 43A (arabinan-specific; EC 3.2.1.-), endo-alpha-L-arabinanase as well as arabinoxylan arabinofuranohydrolase (AXH) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. The GH43_XylA-like subgroup includes Clostridium stercorarium alpha-L-arabinofuranosidase XylA, and enzymes that have been annotated as having beta-xylosidase (EC 3.2.1.37), alpha-L-arabinofuranosidase (EC 3.2.1.55), endo-alpha-L-arabinanase (EC 3.2.1.-) as well as arabinoxylan arabinofuranohydrolase (AXH) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. AXHs specifically hydrolyze the glycosidic bond between arabinofuranosyl substituents and xylopyranosyl backbone residues of arabinoxylan. |
| cd08990 | GH43_AXH_like | 1.52e-45 | 37 | 337 | 2 | 259 | Glycosyl hydrolase family 43 protein, includes arabinoxylan arabinofuranohydrolase, beta-xylosidase, endo-1,4-beta-xylanase, and alpha-L-arabinofuranosidase. This subgroup includes Bacillus subtilis arabinoxylan arabinofuranohydrolase (XynD;BsAXH-m23;BSU18160), Butyrivibrio proteoclasticus alpha-L-arabinofuranosidase (Xsa43E;bpr_I2319), Clostridium stercorarium alpha-L-arabinofuranosidase XylA, and metagenomic beta-xylosidase (EC 3.2.1.37) / alpha-L-arabinofuranosidase (EC 3.2.1.55) CoXyl43. It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. The GH43_AXH-like subgroup includes enzymes that have been characterized with beta-xylosidase, alpha-L-arabinofuranosidase, endo-alpha-L-arabinanase as well as arabinoxylan arabinofuranohydrolase (AXH) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. AXHs specifically hydrolyze the glycosidic bond between arabinofuranosyl substituents and xylopyranosyl backbone residues of arabinoxylan. Metagenomic beta-xylosidase/alpha-L-arabinofuranosidase CoXyl43 shows synergy with Trichoderma reesei cellulases and promotes plant biomass saccharification by degrading xylo-oligosaccharides, such as xylobiose and xylotriose, into the monosaccharide xylose. Studies show that the hydrolytic activity of CoXyl43 is stimulated in the presence of calcium. Several of these enzymes also contain carbohydrate binding modules (CBMs) that bind cellulose or xylan. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd09003 | GH43_XynD-like | 6.72e-33 | 53 | 323 | 20 | 284 | Glycosyl hydrolase family 43 protein such as Bacillus subtilis arabinoxylan arabinofuranohydrolase (XynD;BsAXH-m23;BSU18160). This glycosyl hydrolase family 43 (GH43) subgroup includes characterized Bacillus subtilis arabinoxylan arabinofuranohydrolase (AXH), Caldicellulosiruptor sp. Tok7B.1 beta-1,4-xylanase (EC 3.2.1.8) / alpha-L-arabinosidase (EC 3.2.1.55) XynA, Caldicellulosiruptor sp. Rt69B.1 xylanase C (EC 3.2.1.8) XynC, and Caldicellulosiruptor saccharolyticus beta-xylosidase (EC 3.2.1.37)/ alpha-L-arabinofuranosidase (EC 3.2.1.55) XynF. It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. It belongs to the GH43_AXH-like subgroup which includes enzymes that have been annotated as having beta-xylosidase, alpha-L-arabinofuranosidase and arabinoxylan alpha-L-1,3-arabinofuranohydrolase, xylanase (endo-alpha-L-arabinanase) as well as AXH activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. AXHs specifically hydrolyze the glycosidic bond between arabinofuranosyl substituents and xylopyranosyl backbone residues of arabinoxylan. Bacillus subtilis AXH (BsAXH-m2,3) has been shown to cleave arabinose units from O-2- or O-3-mono-substituted xylose residues and superposition of its structure with known structures of the GH43 exo-acting enzymes, beta-xylosidase and alpha-L-arabinanase, each in complex with their substrate, reveals a different orientation of the sugar backbone. Several of these enzymes also contain carbohydrate binding modules (CBMs) that bind cellulose or xylan. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd18619 | GH43_CoXyl43_like | 7.93e-32 | 53 | 320 | 19 | 284 | Glycosyl hydrolase family 43 protein such as metagenomic beta-xylosidase/alpha-L-arabinofuranosidase CoXyl43. This glycosyl hydrolase family 43 (GH43) subgroup belongs to the GH43_AXH-like subgroup which includes enzymes that have been characterized with beta-xylosidase (EC 3.2.1.37), alpha-L-arabinofuranosidase (EC 3.2.1.55), alpha-1,2-L-arabinofuranosidase 43A (arabinan-specific; EC 3.2.1.-), endo-alpha-L-arabinanase as well as arabinoxylan arabinofuranohydrolase (AXH) activities. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. Included in this subfamily is the metagenomic beta-xylosidase/alpha-L-arabinofuranosidase CoXyl43, which shows synergy with Trichoderma reesei cellulases and promotes plant biomass saccharification by degrading xylo-oligosaccharides, such as xylobiose and xylotriose, into the monosaccharide xylose. Studies show that the hydrolytic activity of CoXyl43 is stimulated in the presence of calcium. Several of these enzymes also contain carbohydrate binding modules (CBMs) that bind cellulose or xylan. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd09004 | GH43_bXyl-like | 1.11e-30 | 53 | 338 | 11 | 256 | Glycosyl hydrolase family 43 protein such as Bacteroides thetaiotaomicron VPI-5482 alpha-L-arabinofuranosidases (BT3675;BT_3675) and (BT3662;BT_3662); includes mostly xylanases. This glycosyl hydrolase family 43 (GH43) subgroup includes enzymes that have been annotated as xylan-digesting beta-xylosidase (EC 3.2.1.37) and xylanase (endo-alpha-L-arabinanase, EC 3.2.1.8) activities, as well the Bacteroides thetaiotaomicron VPI-5482 alpha-L-arabinofuranosidases (EC 3.2.1.55) (BT3675;BT_3675) and (BT3662;BT_3662). It belongs to the glycosyl hydrolase clan F (according to carbohydrate-active enzymes database (CAZY)) which includes family 43 (GH43) and 62 (GH62) families. GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD35875.1 | 2.59e-219 | 11 | 533 | 17 | 540 |
| ADE83588.1 | 2.10e-206 | 23 | 533 | 39 | 551 |
| AHF26057.1 | 4.85e-206 | 1 | 532 | 1 | 547 |
| QVJ80144.1 | 4.85e-205 | 23 | 533 | 39 | 551 |
| BBK86497.1 | 3.71e-198 | 1 | 530 | 1 | 532 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5A8C_A | 2.04e-20 | 53 | 350 | 48 | 315 | ChainA, CARBOHYDRATE BINDING FAMILY 6 [Acetivibrio thermocellus],5A8D_A Chain A, CARBOHYDRATE BINDING FAMILY 6 [Acetivibrio thermocellus] |
| 6XN0_A | 6.85e-18 | 54 | 337 | 60 | 342 | ChainA, Xylosidase [Xanthomonas citri pv. citri str. 306],6XN0_B Chain B, Xylosidase [Xanthomonas citri pv. citri str. 306],6XN1_A Chain A, Xylosidase [Xanthomonas citri pv. citri str. 306],6XN1_B Chain B, Xylosidase [Xanthomonas citri pv. citri str. 306],6XN2_A Chain A, Xylosidase [Xanthomonas citri pv. citri str. 306],6XN2_B Chain B, Xylosidase [Xanthomonas citri pv. citri str. 306] |
| 4MLG_A | 6.76e-16 | 37 | 320 | 15 | 289 | Structureof RS223-Beta-xylosidase [uncultured organism],4MLG_B Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_C Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_D Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_E Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_F Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_G Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_H Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_I Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_J Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_K Structure of RS223-Beta-xylosidase [uncultured organism],4MLG_L Structure of RS223-Beta-xylosidase [uncultured organism] |
| 5GLK_A | 2.81e-14 | 53 | 320 | 41 | 309 | Crystalstructure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compost microbial metagenome, calcium-free form. [uncultured bacterium],5GLK_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compost microbial metagenome, calcium-free form. [uncultured bacterium],5GLL_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome, calcium-bound form [uncultured bacterium],5GLL_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome, calcium-bound form [uncultured bacterium],5GLM_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compost microbial metagenome in complex with xylotriose, calcium-free form. [uncultured bacterium],5GLM_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compost microbial metagenome in complex with xylotriose, calcium-free form. [uncultured bacterium],5GLN_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with xylotriose, calcium-bound form [uncultured bacterium],5GLN_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with xylotriose, calcium-bound form [uncultured bacterium],5GLO_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose, calcium-free form [uncultured bacterium],5GLO_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose, calcium-free form [uncultured bacterium],5GLP_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose, calcium-bound form [uncultured bacterium],5GLP_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose, calcium-bound form [uncultured bacterium],5GLQ_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose and xylotriose, calcium-free form [uncultured bacterium],5GLQ_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose and xylotriose, calcium-free form [uncultured bacterium],5GLR_A Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose and xylotriose, calcium-bound form [uncultured bacterium],5GLR_B Crystal structure of CoXyl43, GH43 beta-xylosidase/alpha-arabinofuranosidase from a compostmicrobial metagenome in complex with l-arabinose and xylotriose, calcium-bound form [uncultured bacterium] |
| 4NOV_A | 5.22e-13 | 37 | 320 | 56 | 305 | Xsa43E,a GH43 family enzyme from Butyrivibrio proteoclasticus [Butyrivibrio proteoclasticus B316] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48790 | 2.94e-50 | 26 | 530 | 7 | 468 | Xylosidase/arabinosidase OS=Thermoclostridium stercorarium OX=1510 GN=xylA PE=1 SV=1 |
| P49943 | 3.75e-15 | 53 | 320 | 25 | 290 | Xylosidase/arabinosidase OS=Bacteroides ovatus OX=28116 GN=xsa PE=2 SV=1 |
| P48791 | 4.71e-15 | 41 | 338 | 11 | 308 | Beta-xylosidase OS=Prevotella ruminicola OX=839 GN=xynB PE=3 SV=1 |
| P45796 | 4.16e-12 | 37 | 532 | 56 | 507 | Arabinoxylan arabinofuranohydrolase OS=Paenibacillus polymyxa OX=1406 GN=xynD PE=1 SV=1 |
| Q45071 | 1.16e-08 | 53 | 322 | 59 | 333 | Arabinoxylan arabinofuranohydrolase OS=Bacillus subtilis (strain 168) OX=224308 GN=xynD PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
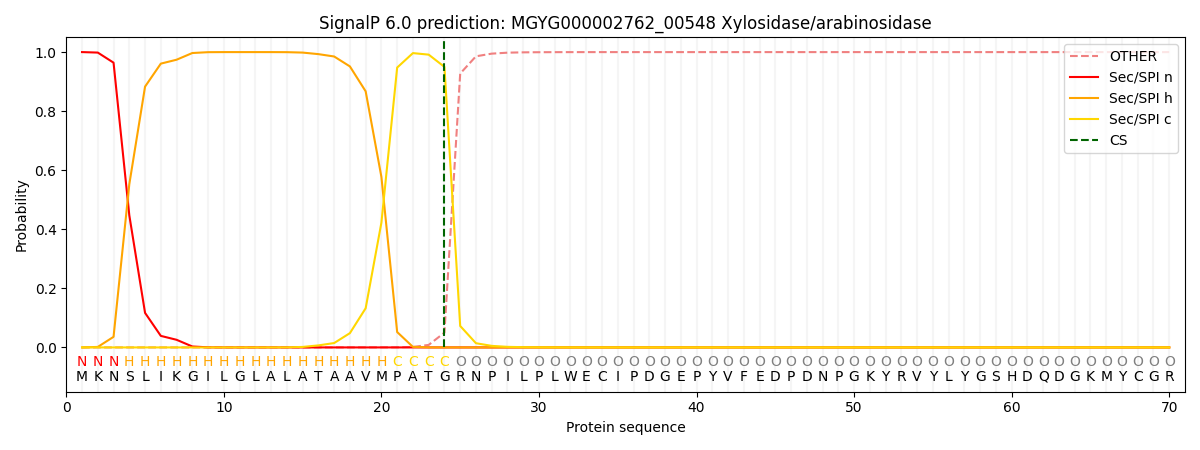
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000420 | 0.998813 | 0.000230 | 0.000201 | 0.000172 | 0.000163 |
