You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002982_00245
You are here: Home > Sequence: MGYG000002982_00245
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
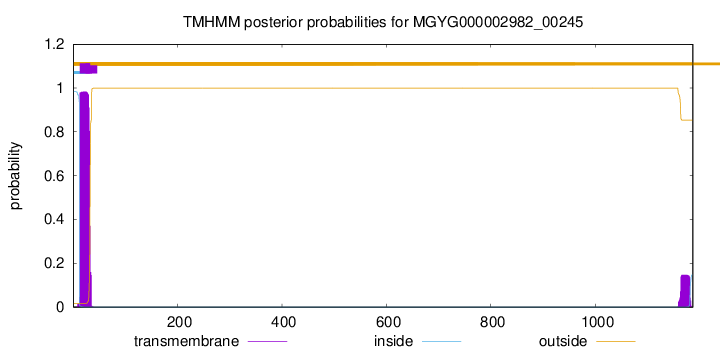
TMHMM annotations
Basic Information help
| Species | Streptococcus rubneri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus rubneri | |||||||||||
| CAZyme ID | MGYG000002982_00245 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | Beta-N-acetylhexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2720; End: 6283 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH20 | 139 | 489 | 6.4e-54 | 0.9287833827893175 |
| GH20 | 582 | 920 | 2.9e-52 | 0.9287833827893175 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3525 | Chb | 0.0 | 132 | 914 | 9 | 732 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
| cd06564 | GH20_DspB_LnbB-like | 4.00e-105 | 578 | 925 | 1 | 326 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| cd06564 | GH20_DspB_LnbB-like | 6.11e-100 | 135 | 494 | 1 | 326 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| COG3525 | Chb | 9.49e-36 | 575 | 960 | 9 | 384 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
| COG3525 | Chb | 2.05e-33 | 105 | 483 | 391 | 732 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AYF93380.1 | 0.0 | 1 | 1187 | 1 | 1187 |
| VED66574.1 | 0.0 | 1 | 1187 | 1 | 1188 |
| VEE18570.1 | 0.0 | 1 | 1187 | 1 | 1188 |
| AGY37538.1 | 0.0 | 1 | 1187 | 1 | 1186 |
| SQH65722.1 | 0.0 | 1 | 1187 | 1 | 1186 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YL5_A | 2.00e-246 | 574 | 1010 | 5 | 442 | Inhibitionof the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_B Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_C Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_D Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],4AZC_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZC_C Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZC_D Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 4AZG_A | 2.07e-246 | 574 | 1010 | 6 | 443 | Differentialinhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZG_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_A Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_C Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_D Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 2YLA_A | 5.65e-246 | 574 | 1010 | 5 | 442 | InhibitionOf The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YLA_B Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YLA_C Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YLA_D Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4] |
| 4AZC_A | 1.60e-245 | 574 | 1010 | 5 | 442 | Differentialinhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 2YL9_A | 1.57e-244 | 574 | 1007 | 22 | 456 | InhibitionOf The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YL9_B Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YL9_C Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4],2YL9_D Inhibition Of The Pneumococcal Virulence Factor Strh And Molecular Insights Into N-Glycan Recognition And Hydrolysis [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49610 | 0.0 | 5 | 1080 | 6 | 1136 | Beta-N-acetylhexosaminidase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=strH PE=1 SV=2 |
| B2UPR7 | 1.65e-06 | 584 | 900 | 239 | 538 | Beta-hexosaminidase Amuc_2136 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_2136 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
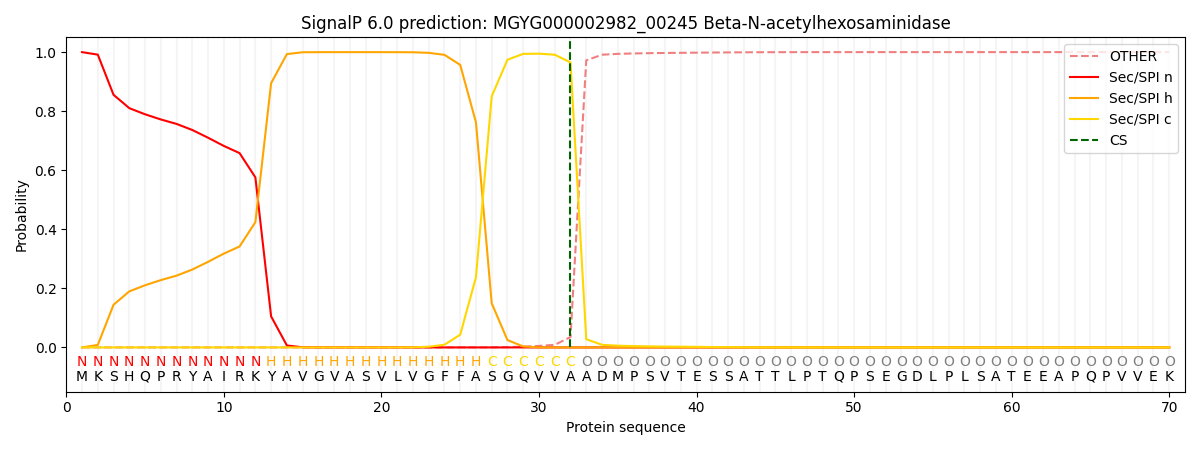
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000666 | 0.998354 | 0.000381 | 0.000208 | 0.000193 | 0.000157 |