You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003113_00819
You are here: Home > Sequence: MGYG000003113_00819
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pauljensenia sp001838165 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Pauljensenia; Pauljensenia sp001838165 | |||||||||||
| CAZyme ID | MGYG000003113_00819 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 142423; End: 143577 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT1 | 47 | 383 | 3.2e-25 | 0.900523560209424 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1819 | YjiC | 2.69e-22 | 1 | 384 | 2 | 400 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| cd03784 | GT1_Gtf-like | 6.28e-22 | 1 | 382 | 1 | 404 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| TIGR00661 | MJ1255 | 2.39e-06 | 120 | 331 | 94 | 294 | conserved hypothetical protein. This model represents nearly the full length of MJ1255 from Methanococcus jannaschii and of an unpublished protein from Vibrio cholerae, as well as the C-terminal half of a protein from Methanobacterium thermoautotrophicum. A small region (~50 amino acids) within the domain appears related to a family of sugar transferases. [Hypothetical proteins, Conserved] |
| COG4671 | COG4671 | 7.16e-05 | 204 | 312 | 195 | 321 | Predicted glycosyl transferase [General function prediction only]. |
| pfam13528 | Glyco_trans_1_3 | 0.003 | 119 | 336 | 95 | 305 | Glycosyl transferase family 1. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCT36430.1 | 3.59e-232 | 8 | 384 | 1 | 377 |
| QGS11935.1 | 4.87e-230 | 8 | 383 | 1 | 376 |
| AHF25143.1 | 2.78e-121 | 1 | 384 | 13 | 406 |
| BAR06867.1 | 4.41e-96 | 1 | 383 | 1 | 383 |
| BAR05900.1 | 1.85e-92 | 1 | 383 | 29 | 416 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3OTG_A | 5.00e-11 | 119 | 377 | 130 | 401 | CrystalStructure of CalG1, Calicheamicin Glycostyltransferase, TDP bound form [Micromonospora echinospora],3OTH_A Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora],3OTH_B Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora] |
| 4RIE_A | 1.42e-09 | 24 | 383 | 20 | 373 | LandomycinGlycosyltransferase LanGT2 [Streptomyces cyanogenus],4RIE_B Landomycin Glycosyltransferase LanGT2 [Streptomyces cyanogenus],4RIF_A Landomycin Glycosyltransferase LanGT2, carbasugar substrate complex [Streptomyces cyanogenus],4RIF_B Landomycin Glycosyltransferase LanGT2, carbasugar substrate complex [Streptomyces cyanogenus] |
| 4RIG_A | 7.97e-09 | 24 | 383 | 20 | 373 | ChimericGlycosyltransferase LanGT2S8Ac [Streptomyces fradiae],4RIG_B Chimeric Glycosyltransferase LanGT2S8Ac [Streptomyces fradiae],4RIH_A Chimeric Glycosyltransferase LanGT2S8Ac, carbasugar substrate complex [Streptomyces cyanogenus],4RIH_B Chimeric Glycosyltransferase LanGT2S8Ac, carbasugar substrate complex [Streptomyces cyanogenus],4RII_A Chimeric Glycosyltransferase LanGT2S8Ac, TDP complex [Streptomyces cyanogenus],4RII_B Chimeric Glycosyltransferase LanGT2S8Ac, TDP complex [Streptomyces cyanogenus] |
| 6KQW_A | 1.93e-08 | 281 | 380 | 282 | 382 | ChainA, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168] |
| 6KQX_A | 1.95e-08 | 281 | 380 | 282 | 382 | ChainA, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168],7BOV_A Chain A, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34539 | 1.07e-07 | 281 | 380 | 282 | 382 | NDP-glycosyltransferase YjiC OS=Bacillus subtilis (strain 168) OX=224308 GN=yjiC PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
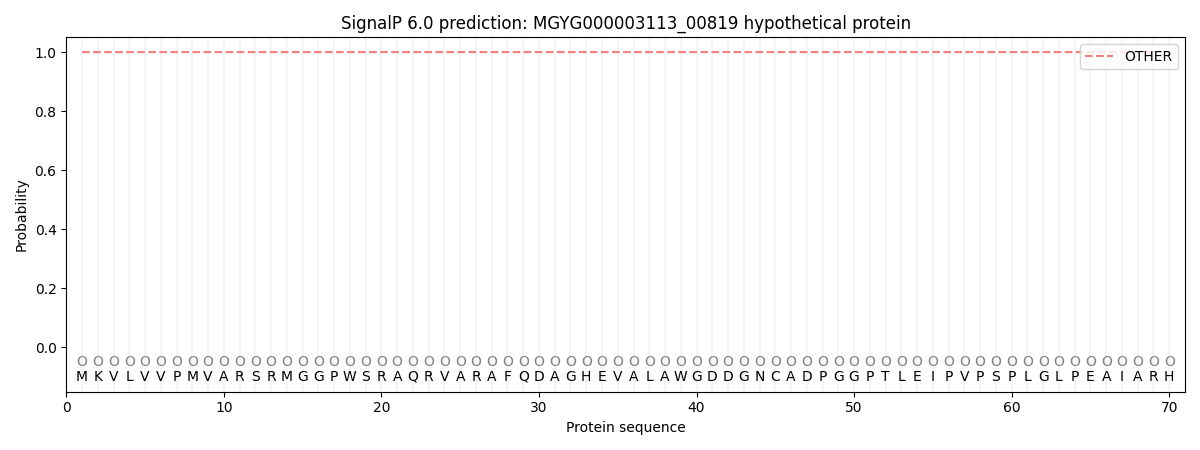
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999995 | 0.000021 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
