You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003113_01751
You are here: Home > Sequence: MGYG000003113_01751
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pauljensenia sp001838165 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Pauljensenia; Pauljensenia sp001838165 | |||||||||||
| CAZyme ID | MGYG000003113_01751 | |||||||||||
| CAZy Family | CE14 | |||||||||||
| CAZyme Description | Mycothiol S-conjugate amidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 15823; End: 16518 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE14 | 6 | 114 | 2.5e-26 | 0.9596774193548387 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2120 | LmbE | 3.04e-36 | 3 | 227 | 12 | 230 | N-acetylglucosaminyl deacetylase, LmbE family [Carbohydrate transport and metabolism]. |
| pfam02585 | PIG-L | 1.65e-30 | 9 | 133 | 4 | 125 | GlcNAc-PI de-N-acetylase. Members of this family are related to PIG-L an N-acetylglucosaminylphosphatidylinositol de-N-acetylase (EC:3.5.1.89) that catalyzes the second step in GPI biosynthesis. |
| PRK02122 | PRK02122 | 0.010 | 10 | 48 | 378 | 416 | glucosamine-6-phosphate deaminase-like protein; Validated |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABK52148.1 | 6.91e-58 | 1 | 217 | 11 | 231 |
| ACM05060.1 | 4.99e-57 | 5 | 230 | 11 | 244 |
| SBO92168.1 | 1.40e-56 | 1 | 206 | 7 | 212 |
| AQZ66804.1 | 2.80e-56 | 1 | 206 | 7 | 212 |
| CCG05052.1 | 3.45e-56 | 1 | 217 | 15 | 234 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5BMO_A | 8.27e-41 | 4 | 191 | 18 | 203 | LnmXprotein, a putative GlcNAc-PI de-N-acetylase from Streptomyces atroolivaceus [Streptomyces atroolivaceus],5BMO_B LnmX protein, a putative GlcNAc-PI de-N-acetylase from Streptomyces atroolivaceus [Streptomyces atroolivaceus],5BMO_C LnmX protein, a putative GlcNAc-PI de-N-acetylase from Streptomyces atroolivaceus [Streptomyces atroolivaceus] |
| 6P2T_A | 1.02e-11 | 4 | 195 | 26 | 201 | BshBfrom Bacillus subtilis complexed with citrate [Bacillus subtilis subsp. subtilis str. 168],6ULL_A BshB from Bacillus subtilis complexed with a substrate analogue [Bacillus subtilis subsp. subtilis str. 168] |
| 2IXD_A | 3.12e-11 | 4 | 195 | 6 | 179 | Crystalstructure of the putative deacetylase BC1534 from Bacillus cereus [Bacillus cereus],2IXD_B Crystal structure of the putative deacetylase BC1534 from Bacillus cereus [Bacillus cereus] |
| 4EWL_A | 1.33e-07 | 4 | 125 | 7 | 147 | ChainA, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase [Mycobacterium tuberculosis],4EWL_B Chain B, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase [Mycobacterium tuberculosis] |
| 1Q74_A | 1.34e-07 | 4 | 125 | 7 | 147 | ChainA, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside Deacetylase (MshB) [Mycobacterium tuberculosis],1Q74_B Chain B, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside Deacetylase (MshB) [Mycobacterium tuberculosis],1Q74_C Chain C, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside Deacetylase (MshB) [Mycobacterium tuberculosis],1Q74_D Chain D, 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside Deacetylase (MshB) [Mycobacterium tuberculosis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0R2W9 | 7.35e-14 | 4 | 149 | 6 | 174 | Mycothiol S-conjugate amidase OS=Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) OX=246196 GN=mca PE=1 SV=1 |
| P9WJN0 | 1.90e-13 | 4 | 167 | 6 | 179 | Mycothiol S-conjugate amidase OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=mca PE=3 SV=1 |
| P9WJN1 | 1.90e-13 | 4 | 167 | 6 | 179 | Mycothiol S-conjugate amidase OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=mca PE=1 SV=1 |
| C1A2R3 | 7.08e-13 | 4 | 194 | 8 | 243 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Rhodococcus erythropolis (strain PR4 / NBRC 100887) OX=234621 GN=mshB PE=3 SV=1 |
| D6Y7M2 | 1.71e-11 | 4 | 168 | 6 | 183 | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase OS=Thermobispora bispora (strain ATCC 19993 / DSM 43833 / CBS 139.67 / JCM 10125 / KCTC 9307 / NBRC 14880 / R51) OX=469371 GN=mshB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
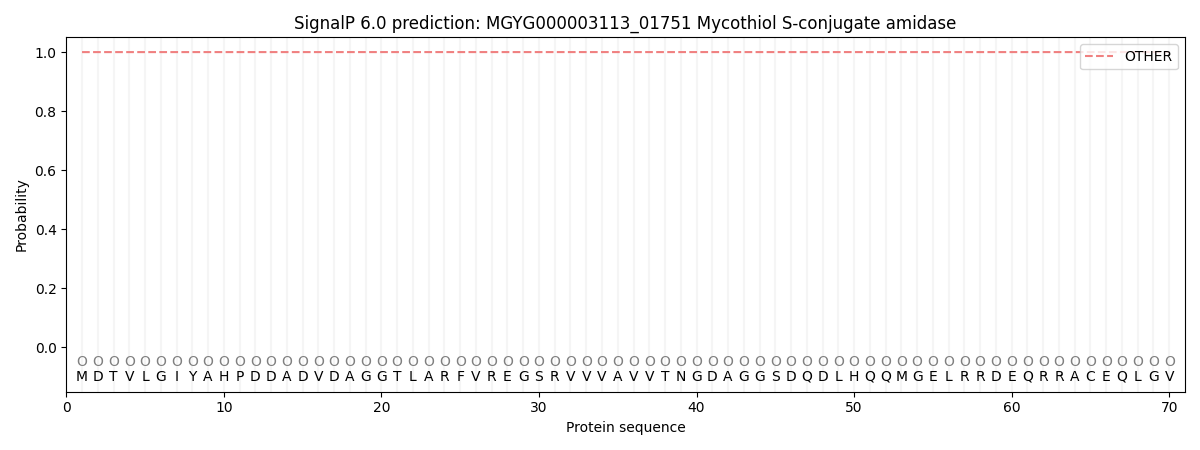
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000057 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
