You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003203_03149
You are here: Home > Sequence: MGYG000003203_03149
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Butyricimonas sp900759925 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Marinifilaceae; Butyricimonas; Butyricimonas sp900759925 | |||||||||||
| CAZyme ID | MGYG000003203_03149 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase D | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11011; End: 12591 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 137 | 276 | 3.8e-22 | 0.9111111111111111 |
| CBM50 | 479 | 524 | 5.2e-16 | 0.95 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16894 | MltD-like | 2.83e-64 | 145 | 273 | 1 | 129 | Membrane-bound lytic murein transglycosylase D and similar proteins. Lytic transglycosylases (LT) catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc). Membrane-bound lytic murein transglycosylase D protein (MltD) family members may have one or more small LysM domains, which may contribute to peptidoglycan binding. Unlike the similar "goose-type" lysozymes, LTs also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| PRK10783 | mltD | 3.87e-63 | 101 | 522 | 77 | 444 | membrane-bound lytic murein transglycosylase D; Provisional |
| pfam01464 | SLT | 6.71e-31 | 140 | 248 | 1 | 110 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| PRK06347 | PRK06347 | 5.85e-19 | 388 | 525 | 473 | 592 | 1,4-beta-N-acetylmuramoylhydrolase. |
| cd13401 | Slt70-like | 8.70e-18 | 136 | 265 | 5 | 139 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRO49268.1 | 0.0 | 1 | 526 | 1 | 525 |
| AZS28203.1 | 0.0 | 1 | 526 | 1 | 525 |
| ADY31171.1 | 7.03e-237 | 43 | 526 | 41 | 524 |
| SNV24466.1 | 7.03e-237 | 43 | 526 | 41 | 524 |
| BAX79662.1 | 6.11e-197 | 15 | 525 | 53 | 563 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2MKX_A | 6.97e-08 | 398 | 439 | 7 | 48 | Solutionstructure of LysM the peptidoglycan binding domain of autolysin AtlA from Enterococcus faecalis [Enterococcus faecalis V583] |
| 1QSA_A | 8.00e-07 | 137 | 277 | 453 | 600 | CrystalStructure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.65 Angstroms Resolution [Escherichia coli],1QTE_A Crystal Structure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.90 A Resolution In Complex With A 1,6- Anhydromurotripeptide [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AEZ8 | 3.62e-41 | 109 | 522 | 74 | 440 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=mltD PE=3 SV=1 |
| P0AEZ7 | 3.62e-41 | 109 | 522 | 74 | 440 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli (strain K12) OX=83333 GN=mltD PE=1 SV=1 |
| P32820 | 7.83e-16 | 134 | 249 | 23 | 138 | Putative tributyltin chloride resistance protein OS=Alteromonas sp. (strain M-1) OX=29457 GN=tbtA PE=3 SV=1 |
| P37710 | 3.14e-11 | 398 | 525 | 363 | 473 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| Q5HRU2 | 1.91e-08 | 376 | 524 | 64 | 190 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=sle1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
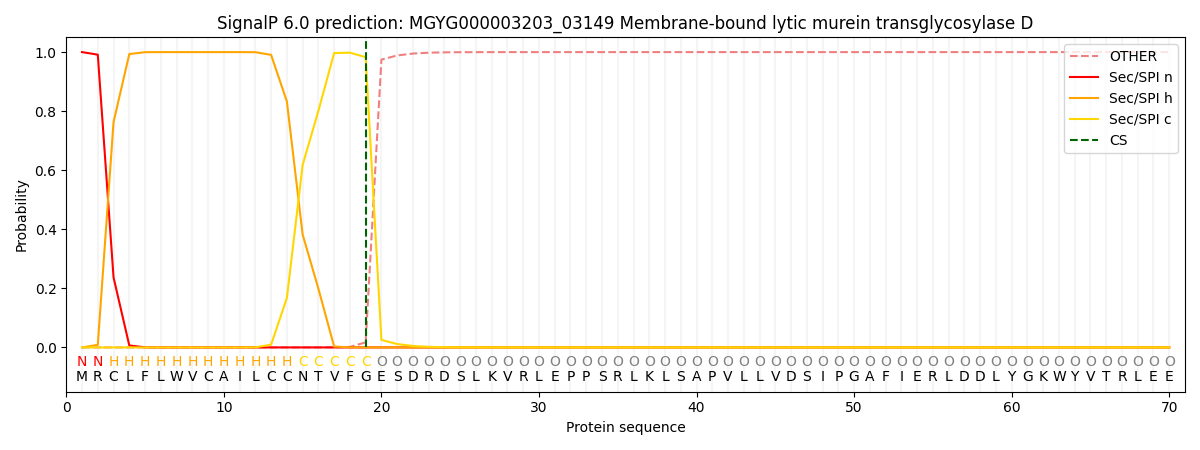
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000210 | 0.999227 | 0.000156 | 0.000145 | 0.000135 | 0.000128 |
