You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003282_00087
You are here: Home > Sequence: MGYG000003282_00087
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coprobacter sp900545915 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Coprobacteraceae; Coprobacter; Coprobacter sp900545915 | |||||||||||
| CAZyme ID | MGYG000003282_00087 | |||||||||||
| CAZy Family | GH85 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 116658; End: 120449 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH85 | 109 | 418 | 3e-54 | 0.946031746031746 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4724 | COG4724 | 6.25e-55 | 41 | 584 | 32 | 553 | Endo-beta-N-acetylglucosaminidase D [Carbohydrate transport and metabolism]. |
| cd06547 | GH85_ENGase | 3.26e-36 | 109 | 450 | 21 | 339 | Endo-beta-N-acetylglucosaminidase (ENGase) hydrolyzes the N-N'-diacetylchitobiosyl core of N-glycosylproteins. The beta-1,4-glycosyl bond located between two N-acetylglucosamine residues is hydrolyzed such that N-acetylglucosamine 1 remains with the protein and N-acetylglucosamine 2 forms the reducing end of the released glycan. ENGase is a key enzyme in the processing of free oligosaccharides in the cytosol of eukaryotes. Oligosaccharides formed in the lumen of the endoplasmic reticulum are transported into the cytosol where they are catabolized by cytosolic ENGases and other enzymes, possibly to maximize the reutilization of the component sugars. ENGases have an eight-stranded alpha/beta barrel topology and are classified as a family 85 glycosyl hydrolase (GH85) domain. The GH85 ENGases are sequence-similar to the family 18 glycosyl hydrolases, also known as GH18 chitinases. An ENGase-like protein is also found in bacteria and is included in this alignment model. |
| pfam13385 | Laminin_G_3 | 2.04e-17 | 949 | 1033 | 63 | 150 | Concanavalin A-like lectin/glucanases superfamily. This domain belongs to the Concanavalin A-like lectin/glucanases superfamily. |
| TIGR04183 | Por_Secre_tail | 1.22e-06 | 1189 | 1263 | 1 | 71 | Por secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber average twenty or more copies of a C-terminal domain, represented by this model, associated with sorting to the outer membrane and covalent modification. |
| pfam18962 | Por_Secre_tail | 3.80e-06 | 1189 | 1256 | 1 | 66 | Secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber have on average twenty or more copies of this C-terminal domain, associated with sorting to the outer membrane and covalent modification. This domain targets proteins to type IX secretion systems and is secreted then cleaved off by a C-terminal signal peptidease. Based on similarity to other families it is likely that this domain adopts an immunoglobulin like fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AHF12130.1 | 0.0 | 1 | 1262 | 1 | 1285 |
| ADK97378.1 | 1.10e-297 | 7 | 1263 | 6 | 1256 |
| ASE18174.1 | 1.10e-297 | 7 | 1263 | 6 | 1256 |
| EFC71170.1 | 6.90e-294 | 8 | 1263 | 4 | 1249 |
| EFC71169.1 | 8.57e-294 | 10 | 1263 | 6 | 1256 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2VTF_A | 1.78e-42 | 34 | 686 | 8 | 608 | X-raycrystal structure of the Endo-beta-N-acetylglucosaminidase from Arthrobacter protophormiae E173Q mutant reveals a TIM barrel catalytic domain and two ancillary domains [Glutamicibacter protophormiae],2VTF_B X-ray crystal structure of the Endo-beta-N-acetylglucosaminidase from Arthrobacter protophormiae E173Q mutant reveals a TIM barrel catalytic domain and two ancillary domains [Glutamicibacter protophormiae] |
| 3FHA_A | 9.67e-42 | 34 | 686 | 3 | 603 | ChainA, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHA_B Chain B, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHA_C Chain C, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHA_D Chain D, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae] |
| 3FHQ_A | 1.73e-41 | 34 | 686 | 3 | 603 | ChainA, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHQ_B Chain B, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHQ_D Chain D, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae],3FHQ_F Chain F, Endo-beta-N-acetylglucosaminidase [Glutamicibacter protophormiae] |
| 2W91_A | 2.58e-33 | 33 | 686 | 7 | 627 | Structureof a Streptococcus pneumoniae family 85 glycoside hydrolase, Endo-D. [Streptococcus pneumoniae TIGR4],2W92_A Structure of a Streptococcus pneumoniae family 85 glycoside hydrolase, Endo-D, in complex with NAG-thiazoline. [Streptococcus pneumoniae TIGR4] |
| 3GDB_A | 8.56e-33 | 33 | 686 | 158 | 778 | Crystalstructure of Spr0440 glycoside hydrolase domain, Endo-D from Streptococcus pneumoniae R6 [Streptococcus pneumoniae R6] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A1L251 | 2.57e-07 | 115 | 516 | 130 | 489 | Cytosolic endo-beta-N-acetylglucosaminidase OS=Danio rerio OX=7955 GN=engase PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
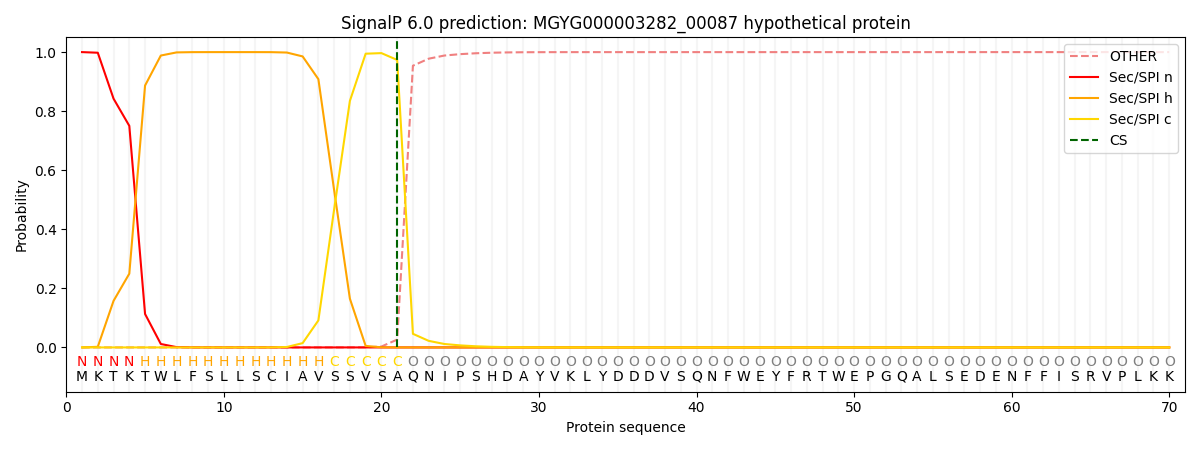
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000389 | 0.998798 | 0.000260 | 0.000173 | 0.000172 | 0.000165 |
