You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003475_00347
You are here: Home > Sequence: MGYG000003475_00347
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Selenomonas_A sp900769615 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Selenomonadales; Selenomonadaceae; Selenomonas_A; Selenomonas_A sp900769615 | |||||||||||
| CAZyme ID | MGYG000003475_00347 | |||||||||||
| CAZy Family | GH53 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 65953; End: 67815 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH53 | 43 | 386 | 3.4e-133 | 0.9941520467836257 |
| CBM61 | 478 | 615 | 2.8e-35 | 0.9787234042553191 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3867 | GanB | 1.40e-146 | 19 | 390 | 16 | 394 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
| pfam07745 | Glyco_hydro_53 | 1.49e-140 | 43 | 387 | 1 | 333 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| pfam07532 | Big_4 | 9.23e-08 | 395 | 462 | 1 | 59 | Bacterial Ig-like domain (group 4). This family consists of bacterial domains with an Ig-like fold. Members of this family are found in a variety of bacterial surface proteins. |
| cd19608 | GH113_mannanase-like | 6.78e-07 | 71 | 365 | 24 | 294 | Glycoside hydrolase family 113 beta-1,4-mannanase and similar proteins. Mannan endo-1,4-beta mannosidase (E.C 3.2.1.78) randomly cleaves (1->4)-beta-D-mannosidic linkages in mannans, galactomannans and glucomannans and is also called beta-1,4-mannanase, endo-1,4-beta-mannanase, endo-beta-1,4-mannase, beta-mannanase B, beta-1, 4-mannan 4-mannanohydrolase, endo-beta-mannanase, beta-D-mannanase, 1,4-beta-D-mannan mannanohydrolase, and 4-beta-D-mannan mannanohydrolase. (1->4)-beta-linked mannans are polysaccharides with a linear polymer backbone of (1->4)-beta-linked mannose units (in plants and fungi) or alternating mannose and glucose/galactose units (glucomannan in plants and fungi, and galactomannan and galactoglucomannan in plants), such as in the hemicellulose fraction of hard- and softwoods. Complete degradation of mannan requires a series of enzymes, including beta-1,4-mannanase. According to the CAZy database beta-1,4-mannanases are grouped into various glycoside hydrolase (GH) families; GH family 113 beta-1,4-mannanases include mostly bacterial and archaeal sequences. |
| cd04080 | CBM6_cellulase-like | 1.93e-04 | 537 | 616 | 67 | 143 | Carbohydrate Binding Module 6 (CBM6); appended to glycoside hydrolase (GH) domains, including GH5 (cellulase). This family includes carbohydrate binding module 6 (CBM6) domains that are appended to several glycoside hydrolase (GH) domains, including GH5 (cellulase) and GH16, as well as to coagulation factor 5/8 carbohydrate-binding domains. CBM6s are non-catalytic carbohydrate binding domains that facilitate the strong binding of the GH catalytic modules with their dedicated, insoluble substrates. The CBM6s are appended to GHs that display a diversity of substrate specificities. For some members of this family information is available about the specific substrates of the appended GH domains. It includes the CBM domains of various enzymes involved in cell wall degradation including, an extracellular beta-1,3-glucanase from Lysobacter enzymogenes encoded by the gluC gene (its catalytic domain belongs to the GH16 family), the tandem CBM domains of Pseudomonas sp. PE2 beta-1,3(4)-glucanase A (its catalytic domain also belongs to GH16), and a family 6 CBM from Cellvibrio mixtus Endoglucanase 5A (CmCBM6) which binds to the beta1,4-beta1,3-mixed linked glucans lichenan, and barley beta-glucan, cello-oligosaccharides, insoluble forms of cellulose, the beta1,3-glucan laminarin, and xylooligosaccharides, and the CBM6 of Fibrobacter succinogenes S85 XynD xylanase, appended to a GH10 domain, and Cellvibrio japonicas Cel5G appended to a GH5 (cellulase) domain. GH5 (cellulase) family includes enzymes with several known activities such as endoglucanase, beta-mannanase, and xylanase, which are involved in the degradation of cellulose and xylans. GH16 family includes enzymes with lichenase, xyloglucan endotransglycosylase (XET), and beta-agarase activities. CBM6 is an unusual CBM as it represents a chimera of two distinct binding sites with different modes of binding: binding site I within the loop regions and binding site II on the concave face of the beta-sandwich fold. For CmCBM6 it has been shown that these two binding sites have different ligand specificities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAL82134.1 | 7.14e-272 | 1 | 619 | 1 | 615 |
| QIB60860.1 | 4.61e-223 | 1 | 619 | 1 | 623 |
| AFK85183.1 | 8.77e-182 | 32 | 620 | 42 | 639 |
| AEI40749.1 | 1.83e-179 | 23 | 617 | 27 | 632 |
| AFC29355.1 | 6.44e-179 | 23 | 617 | 27 | 632 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7OSK_A | 4.08e-101 | 30 | 388 | 38 | 392 | ChainA, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230],7OSK_B Chain B, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230] |
| 1R8L_A | 6.48e-70 | 25 | 388 | 7 | 389 | Thestructure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1R8L_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis [Bacillus licheniformis],1UR0_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR0_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_A The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],1UR4_B The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. [Bacillus licheniformis],2CCR_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2CCR_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_A Structure of Beta-1,4-Galactanase [Bacillus licheniformis],2J74_B Structure of Beta-1,4-Galactanase [Bacillus licheniformis] |
| 2GFT_A | 4.86e-69 | 25 | 388 | 7 | 389 | ChainA, Glycosyl Hydrolase Family 53 [Bacillus licheniformis],2GFT_B Chain B, Glycosyl Hydrolase Family 53 [Bacillus licheniformis] |
| 4BF7_A | 4.10e-53 | 44 | 391 | 22 | 351 | Emericillanidulans endo-beta-1,4-galactanase [Aspergillus nidulans] |
| 1HJS_A | 9.04e-52 | 44 | 388 | 5 | 330 | Structureof two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_A Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48843 | 2.25e-82 | 42 | 388 | 6 | 347 | Uncharacterized protein in bgaB 5'region (Fragment) OS=Niallia circulans OX=1397 PE=3 SV=1 |
| O07013 | 2.00e-70 | 32 | 388 | 43 | 418 | Endo-beta-1,4-galactanase OS=Bacillus subtilis (strain 168) OX=224308 GN=ganB PE=1 SV=1 |
| Q65CX5 | 7.01e-69 | 25 | 388 | 32 | 414 | Endo-beta-1,4-galactanase OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=ganB PE=1 SV=1 |
| A1D3T4 | 1.27e-54 | 44 | 352 | 26 | 323 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=galA PE=3 SV=1 |
| Q0CTQ7 | 1.07e-53 | 44 | 359 | 19 | 323 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=galA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
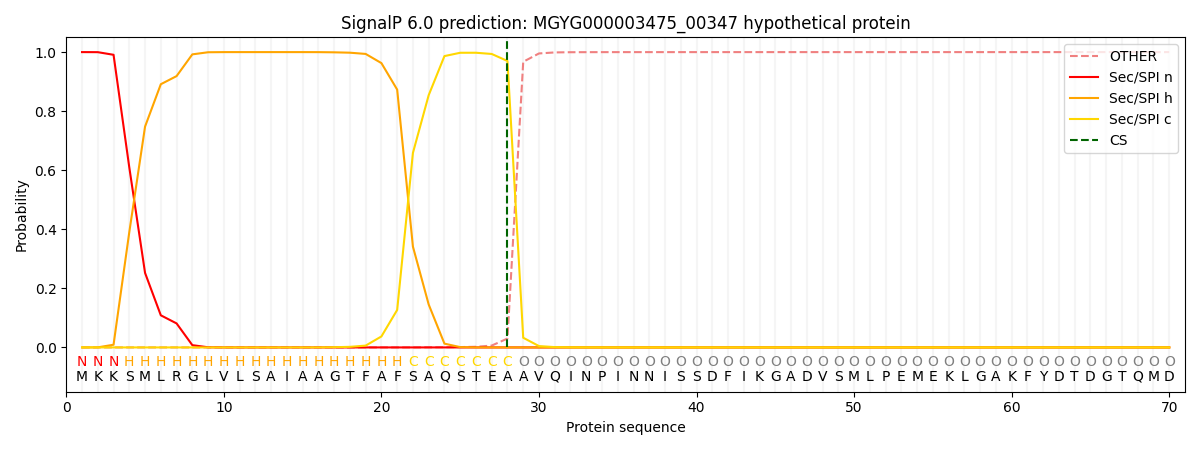
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000229 | 0.999060 | 0.000201 | 0.000181 | 0.000170 | 0.000145 |
