You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003584_00798
You are here: Home > Sequence: MGYG000003584_00798
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | HGM11412 sp900770185 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; Borkfalkiaceae; HGM11412; HGM11412 sp900770185 | |||||||||||
| CAZyme ID | MGYG000003584_00798 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3099; End: 5405 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 402 | 759 | 2.5e-50 | 0.9324324324324325 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06548 | GH18_chitinase | 3.88e-51 | 487 | 755 | 102 | 322 | The GH18 (glycosyl hydrolases, family 18) type II chitinases hydrolyze chitin, an abundant polymer of N-acetylglucosamine and have been identified in bacteria, fungi, insects, plants, viruses, and protozoan parasites. The structure of this domain is an eight-stranded alpha/beta barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. |
| smart00636 | Glyco_18 | 6.57e-51 | 438 | 751 | 31 | 330 | Glyco_18 domain. |
| pfam00704 | Glyco_hydro_18 | 1.47e-47 | 438 | 755 | 28 | 307 | Glycosyl hydrolases family 18. |
| COG3325 | ChiA | 4.41e-44 | 469 | 764 | 128 | 432 | Chitinase, GH18 family [Carbohydrate transport and metabolism]. |
| cd02872 | GH18_chitolectin_chitotriosidase | 1.56e-37 | 469 | 751 | 72 | 337 | This conserved domain family includes a large number of catalytically inactive chitinase-like lectins (chitolectins) including YKL-39, YKL-40 (HCGP39), YM1, oviductin, and AMCase (acidic mammalian chitinase), as well as catalytically active chitotriosidases. The conserved domain is an eight-stranded alpha/beta barrel fold belonging to the family 18 glycosyl hydrolases. The fold has a pronounced active-site cleft at the C-terminal end of the beta-barrel. The chitolectins lack a key active site glutamate (the proton donor required for hydrolytic activity) but retain highly conserved residues involved in oligosaccharide binding. Chitotriosidase is a chitinolytic enzyme expressed in maturing macrophages, which suggests that it plays a part in antimicrobial defense. Chitotriosidase hydrolyzes chitotriose, as well as colloidal chitin to yield chitobiose and is therefore considered an exochitinase. Chitotriosidase occurs in two major forms, the large form being converted to the small form by either RNA or post-translational processing. Although the small form, containing the chitinase domain alone, is sufficient for the chitinolytic activity, the additional C-terminal chitin-binding domain of the large form plays a role in processing colloidal chitin. The chitotriosidase gene is nonessential in humans, as about 35% of the population are heterozygous and 6% homozygous for an inactivated form of the gene. HCGP39 is a 39-kDa human cartilage glycoprotein thought to play a role in connective tissue remodeling and defense against pathogens. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATL45848.1 | 9.87e-34 | 431 | 768 | 54 | 378 |
| ARS42913.1 | 1.52e-33 | 451 | 768 | 57 | 353 |
| QRZ20136.1 | 7.03e-33 | 494 | 768 | 150 | 421 |
| AKD54683.1 | 3.38e-32 | 455 | 768 | 80 | 372 |
| QTH46681.1 | 3.85e-32 | 453 | 768 | 121 | 437 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1ITX_A | 2.29e-29 | 493 | 768 | 146 | 419 | ChainA, Glycosyl Hydrolase [Niallia circulans] |
| 6KXN_A | 3.92e-28 | 495 | 768 | 101 | 370 | Crystalstructure of W50A mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis],6KXN_B Crystal structure of W50A mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis] |
| 6KST_A | 3.92e-28 | 495 | 768 | 101 | 370 | Crystalstructure of the catalytic domain of chitinase ChiL from Chitiniphilus shinanonensis (CsChiL) [Chitiniphilus shinanonensis],6KST_B Crystal structure of the catalytic domain of chitinase ChiL from Chitiniphilus shinanonensis (CsChiL) [Chitiniphilus shinanonensis],6KXL_A Crystal structure of the catalytic domain of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis],6KXL_B Crystal structure of the catalytic domain of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis] |
| 6INX_A | 6.02e-28 | 468 | 768 | 63 | 342 | Structuralinsights into a novel glycoside hydrolase family 18 N-acetylglucosaminidase from Paenibacillus barengoltzii [Paenibacillus barengoltzii] |
| 6KXM_A | 1.75e-27 | 495 | 768 | 101 | 370 | Crystalstructure of D157N mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis],6KXM_B Crystal structure of D157N mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P20533 | 1.28e-27 | 493 | 768 | 178 | 451 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
| Q9W5U2 | 1.71e-20 | 452 | 755 | 1464 | 1753 | Probable chitinase 10 OS=Drosophila melanogaster OX=7227 GN=Cht10 PE=2 SV=2 |
| A6N6J0 | 6.94e-20 | 444 | 755 | 100 | 390 | Endochitinase 46 OS=Trichoderma harzianum OX=5544 GN=chit46 PE=1 SV=1 |
| P48827 | 2.08e-19 | 444 | 761 | 99 | 391 | Endochitinase 42 OS=Trichoderma harzianum OX=5544 GN=chit42 PE=1 SV=1 |
| P11797 | 2.76e-19 | 488 | 755 | 113 | 408 | Chitinase B OS=Serratia marcescens OX=615 GN=chiB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
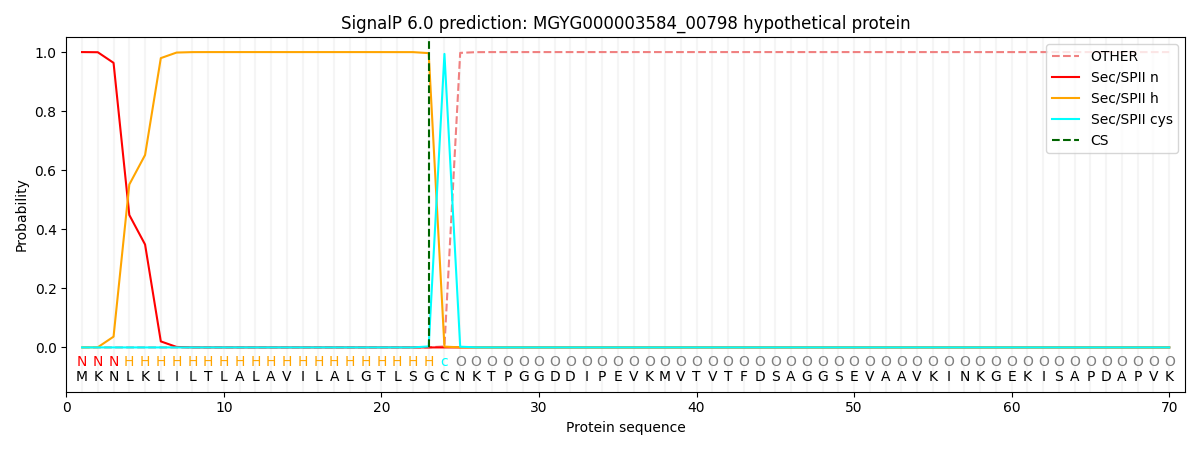
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000053 | 0.000000 | 0.000000 | 0.000000 |
