You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003622_01543
You are here: Home > Sequence: MGYG000003622_01543
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900770845 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900770845 | |||||||||||
| CAZyme ID | MGYG000003622_01543 | |||||||||||
| CAZy Family | GH95 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 16001; End: 24202 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH95 | 1474 | 2196 | 2.6e-206 | 0.9875346260387812 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14498 | Glyco_hyd_65N_2 | 1.24e-45 | 1479 | 1687 | 1 | 232 | Glycosyl hydrolase family 65, N-terminal domain. This domain represents a domain found to the N-terminus of the glycosyl hydrolase 65 family catalytic domain. |
| pfam13402 | Peptidase_M60 | 3.01e-14 | 698 | 970 | 1 | 268 | Peptidase M60, enhancin and enhancin-like. This family of peptidases contains a zinc metallopeptidase motif (HEXXHX(8,28)E) and possesses mucinase activity. It includes the viral enhancins as well as enhancin-like peptidases from bacterial species. Enhancins are a class of metalloproteases found in some baculoviruses that enhance viral infection by degrading the peritrophic membrane (PM) of the insect midgut. Bacterial enhancins are found to be cytotoxic when compared to viral enhancin, however, suggesting that the bacterial enhancins do not enhance infection in the same way as viral enhancin. Bacterial enhancins may have evolved a distinct biochemical function. These bacterial domains are peptidases targetting host glycoproteins and thus probably play an important role in successful colonisation of both vertebrate mucosal surfaces and the invertebrate digestive tract by both mutualistic and pathogenic microbes. This family has been augmented by a merge with the sequences in the Enhancin Pfam family. |
| pfam00754 | F5_F8_type_C | 3.11e-04 | 2256 | 2371 | 6 | 120 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam18630 | Peptidase_M60_C | 0.002 | 1069 | 1140 | 1 | 65 | Peptidase M60 C-terminal domain. This is C-terminal domain (CTD) of M60-peptidases pfam13402. It Can also be found at the C-terminal region of gingipain B (RgpB) from P. gingivalis. It was found to possess a typical Ig-like fold encompassing seven antiparallel beta-strands organized in two beta-sheets, packed into a beta-sandwich structure that can spontaneously dimerize through C-terminal strand swapping. Translocation of gingipains from the periplasm across the OM is dependent on the conserved CTD, which appears to be important for secretion of the proteins and in particular, truncation of the last few C-terminal residues of this domain leads to accumulation of gingipains in the periplasm. Subsequently, the T9SS targeting signal was demonstrated to reside within the last 22 residues at the C-terminus of the CTD. During gingipain translocation across the OM, the CTD is cleaved off by PorU. |
| pfam00754 | F5_F8_type_C | 0.003 | 2404 | 2530 | 6 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBA29879.1 | 0.0 | 1158 | 2253 | 28 | 1126 |
| QUB76594.1 | 0.0 | 1158 | 2245 | 28 | 1119 |
| QUB66762.1 | 0.0 | 1158 | 2290 | 28 | 1140 |
| QUB57135.1 | 0.0 | 1158 | 2253 | 28 | 1126 |
| QUB59130.1 | 0.0 | 1158 | 2253 | 28 | 1126 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2RDY_A | 1.91e-114 | 1481 | 2237 | 7 | 796 | ChainA, BH0842 protein [Halalkalibacterium halodurans C-125],2RDY_B Chain B, BH0842 protein [Halalkalibacterium halodurans C-125] |
| 4UFC_A | 1.54e-110 | 1469 | 2185 | 14 | 736 | Crystalstructure of the GH95 enzyme BACOVA_03438 [Bacteroides ovatus],4UFC_B Crystal structure of the GH95 enzyme BACOVA_03438 [Bacteroides ovatus] |
| 2EAB_A | 5.27e-91 | 1479 | 2228 | 20 | 892 | Crystalstructure of 1,2-a-L-fucosidase from Bifidobacterium bifidum (apo form) [Bifidobacterium bifidum],2EAB_B Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum (apo form) [Bifidobacterium bifidum],2EAC_A Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum in complex with deoxyfuconojirimycin [Bifidobacterium bifidum],2EAC_B Crystal structure of 1,2-a-L-fucosidase from Bifidobacterium bifidum in complex with deoxyfuconojirimycin [Bifidobacterium bifidum] |
| 2EAD_A | 3.15e-90 | 1479 | 2228 | 20 | 892 | ChainA, Alpha-fucosidase [Bifidobacterium bifidum],2EAD_B Chain B, Alpha-fucosidase [Bifidobacterium bifidum] |
| 2EAE_A | 5.61e-90 | 1479 | 2228 | 19 | 891 | ChainA, Alpha-fucosidase [Bifidobacterium bifidum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8L7W8 | 1.02e-89 | 1479 | 2193 | 54 | 815 | Alpha-L-fucosidase 2 OS=Arabidopsis thaliana OX=3702 GN=FUC95A PE=1 SV=1 |
| A2R797 | 1.03e-81 | 1468 | 2173 | 12 | 759 | Probable alpha-fucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=afcA PE=3 SV=1 |
| Q5AU81 | 3.10e-71 | 1495 | 2171 | 46 | 776 | Alpha-fucosidase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=afcA PE=1 SV=1 |
| Q2USL3 | 8.54e-55 | 1467 | 2171 | 8 | 695 | Probable alpha-fucosidase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=afcA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
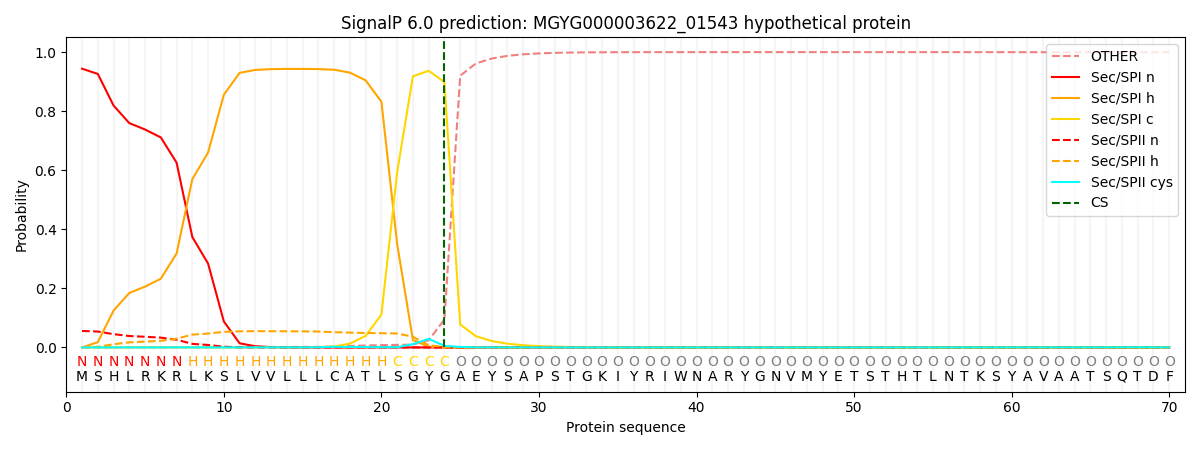
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001506 | 0.938067 | 0.059561 | 0.000295 | 0.000277 | 0.000258 |
