You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003708_01355
You are here: Home > Sequence: MGYG000003708_01355
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
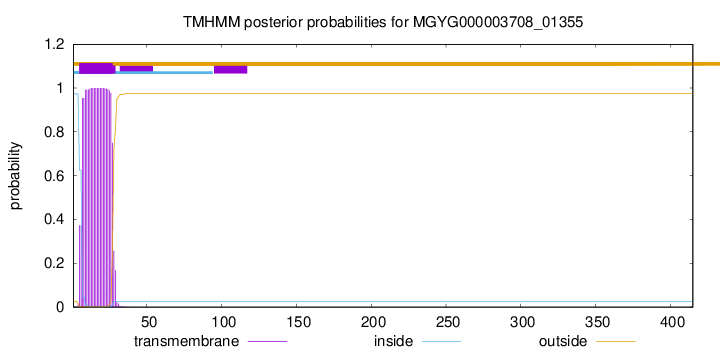
TMHMM annotations
Basic Information help
| Species | Sedimentibacter sp003457065 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Tissierellales; Sedimentibacteraceae; Sedimentibacter; Sedimentibacter sp003457065 | |||||||||||
| CAZyme ID | MGYG000003708_01355 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2303; End: 3550 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 119 | 400 | 1.5e-49 | 0.9527027027027027 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06545 | GH18_3CO4_chitinase | 1.14e-48 | 120 | 400 | 1 | 248 | The Bacteroides thetaiotaomicron protein represented by pdb structure 3CO4 is an uncharacterized bacterial member of the family 18 glycosyl hydrolases with homologs found in Flavobacterium, Stigmatella, and Pseudomonas. |
| pfam00704 | Glyco_hydro_18 | 6.38e-48 | 119 | 394 | 1 | 307 | Glycosyl hydrolases family 18. |
| smart00636 | Glyco_18 | 3.45e-44 | 119 | 365 | 1 | 263 | Glyco_18 domain. |
| cd06548 | GH18_chitinase | 2.55e-34 | 120 | 341 | 1 | 269 | The GH18 (glycosyl hydrolases, family 18) type II chitinases hydrolyze chitin, an abundant polymer of N-acetylglucosamine and have been identified in bacteria, fungi, insects, plants, viruses, and protozoan parasites. The structure of this domain is an eight-stranded alpha/beta barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. |
| cd00598 | GH18_chitinase-like | 3.21e-34 | 120 | 299 | 1 | 179 | The GH18 (glycosyl hydrolase, family 18) type II chitinases hydrolyze chitin, an abundant polymer of beta-1,4-linked N-acetylglucosamine (GlcNAc) which is a major component of the cell wall of fungi and the exoskeleton of arthropods. Chitinases have been identified in viruses, bacteria, fungi, protozoan parasites, insects, and plants. The structure of the GH18 domain is an eight-stranded beta/alpha barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. The GH18 family includes chitotriosidase, chitobiase, hevamine, zymocin-alpha, narbonin, SI-CLP (stabilin-1 interacting chitinase-like protein), IDGF (imaginal disc growth factor), CFLE (cortical fragment-lytic enzyme) spore hydrolase, the type III and type V plant chitinases, the endo-beta-N-acetylglucosaminidases, and the chitolectins. The GH85 (glycosyl hydrolase, family 85) ENGases (endo-beta-N-acetylglucosaminidases) are closely related to the GH18 chitinases and are included in this alignment model. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QOX64831.1 | 3.83e-134 | 117 | 406 | 46 | 334 |
| QAT42625.1 | 8.59e-120 | 116 | 408 | 35 | 332 |
| QIB69208.1 | 4.57e-116 | 117 | 408 | 32 | 328 |
| ARE87297.1 | 1.84e-111 | 116 | 398 | 34 | 316 |
| AOY76825.1 | 1.84e-111 | 116 | 398 | 34 | 316 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6XYZ_A | 4.49e-26 | 112 | 408 | 11 | 303 | Crystalstructure of the GH18 chitinase ChiB from the chitin utilization locus of Flavobacterium johnsoniae [Flavobacterium johnsoniae UW101] |
| 5XWQ_A | 4.78e-19 | 134 | 337 | 15 | 239 | Crystalstructure of chitinase (RmChi1) from Rhizomucor miehei (sp p32 2 1, MR) [Rhizomucor miehei],5YUQ_A Chain A, Chintase [Rhizomucor miehei],5YUQ_B Chain B, Chintase [Rhizomucor miehei],7FBT_A Chain A, Chitinase [Rhizomucor miehei] |
| 3WL1_A | 5.52e-19 | 119 | 335 | 8 | 256 | Crystalstructure of Ostrinia furnacalis Group I chitinase catalytic domain in complex with reaction products (GlcNAc)2,3 [Ostrinia furnacalis],3WQV_A Crystal structure of Ostrinia furnacalis Group I chitinase catalytic domain in complex with a(GlcN)5 [Ostrinia furnacalis],3WQW_A Crystal structure of Ostrinia furnacalis Group I chitinase catalytic domain in complex with a(GlcN)6 [Ostrinia furnacalis] |
| 3W4R_A | 1.13e-18 | 119 | 335 | 26 | 274 | Crystalstructure of an insect chitinase from the Asian corn borer, Ostrinia furnacalis [Ostrinia furnacalis] |
| 3WKZ_A | 1.36e-18 | 119 | 335 | 8 | 256 | CrystalStructure of the Ostrinia furnacalis Group I Chitinase catalytic domain E148Q mutant [Ostrinia furnacalis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9W5U2 | 3.44e-20 | 134 | 335 | 1431 | 1655 | Probable chitinase 10 OS=Drosophila melanogaster OX=7227 GN=Cht10 PE=2 SV=2 |
| Q5AM60 | 4.55e-16 | 134 | 335 | 41 | 264 | Chitinase 4 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=CHT4 PE=3 SV=1 |
| Q873X9 | 1.06e-15 | 130 | 335 | 58 | 301 | Endochitinase B1 OS=Neosartorya fumigata OX=746128 GN=chiB1 PE=1 SV=1 |
| E9QRF2 | 1.06e-15 | 130 | 335 | 58 | 301 | Endochitinase B1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=chiB1 PE=3 SV=1 |
| P36362 | 8.54e-15 | 119 | 335 | 24 | 272 | Endochitinase OS=Manduca sexta OX=7130 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
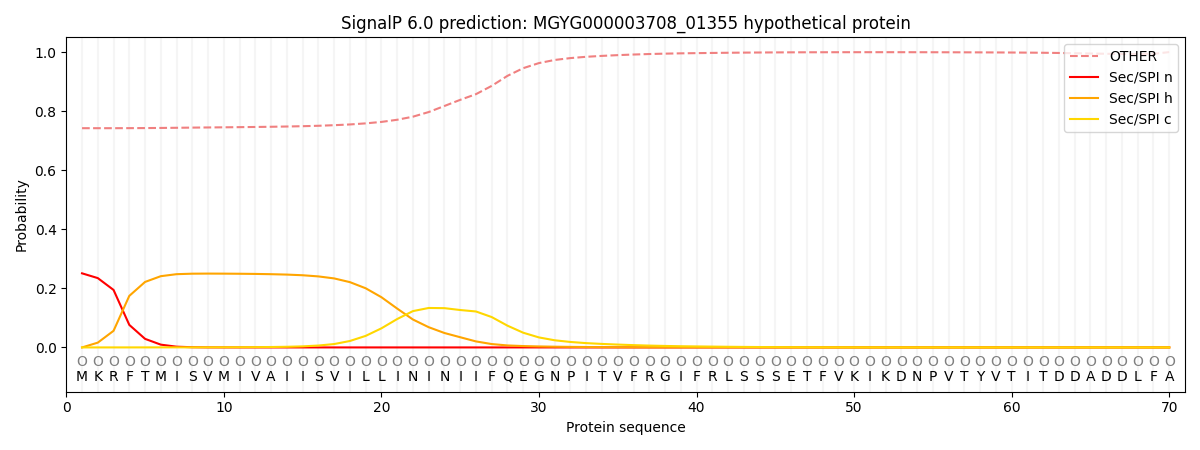
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.755472 | 0.232730 | 0.004150 | 0.001016 | 0.000667 | 0.005973 |