You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003816_00616
You are here: Home > Sequence: MGYG000003816_00616
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; | |||||||||||
| CAZyme ID | MGYG000003816_00616 | |||||||||||
| CAZy Family | GH57 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1395; End: 2825 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH57 | 6 | 322 | 2.4e-57 | 0.8067885117493473 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10795 | GH57N_MJA1_like | 7.59e-167 | 3 | 311 | 1 | 306 | N-terminal catalytic domain of a thermoactive alpha-amylase from Methanococcus jannaschii and similar proteins; glycoside hydrolase family 57 (GH57). The subfamily is represented by a thermostable alpha-amylase (MJA1, EC 3.2.1.1) encoded from the hyperthermophilic archaeon Methanococcus jannaschii locus, M J1611. MJA1 has a broad pH optimum 5.0-8.0. It exhibits extremely thermophilic alpha-amylase activity that catalyzes the hydrolysis of large sugar polymers with alpha-l,6 and alpha-l,4 linkages, and yields products including glucose polymers of 1-7 units. MJ1611 also encodes another alpha-amylase with catalytic features distinct from MJA1, which belongs to glycoside hydrolase family 13 (GH-13), and is not included here. This subfamily also includes many uncharacterized proteins found in bacteria and archaea. |
| pfam03065 | Glyco_hydro_57 | 2.97e-74 | 6 | 293 | 2 | 293 | Glycosyl hydrolase family 57. This family includes alpha-amylase (EC:3.2.1.1), 4--glucanotransferase (EC:2.4.1.-) and amylopullulanase enzymes. |
| COG1449 | COG1449 | 1.20e-66 | 1 | 454 | 85 | 556 | Alpha-amylase/alpha-mannosidase, GH57 family [Carbohydrate transport and metabolism]. |
| cd01022 | GH57N_like | 1.12e-34 | 5 | 310 | 1 | 313 | N-terminal catalytic domain of heat stable retaining glycoside hydrolase family 57. Glycoside hydrolase family 57(GH57) is a chiefly prokaryotic family with the majority of thermostable enzymes coming from extremophiles (many of these are archaeal hyperthermophiles), which exhibit the enzyme specificities of alpha-amylase (EC 3.2.1.1), 4-alpha-glucanotransferase (EC 2.4.1.25), amylopullulanase (EC 3.2.1.1/41), and alpha-galactosidase (EC 3.2.1.22). This family also includes many hypothetical proteins with uncharacterized activity and specificity. GH57s cleave alpha-glycosidic bonds by employing a retaining mechanism, which involves a glycosyl-enzyme intermediate, allowing transglycosylation. |
| cd10793 | GH57N_TLGT_like | 3.51e-16 | 6 | 161 | 2 | 135 | N-terminal catalytic domain of 4-alpha-glucanotransferase; glycoside hydrolase family 57 (GH57). 4-alpha-glucanotransferase (TLGT, EC 2.4.1.25) plays a key role in the maltose metabolism. It catalyzes the disproportionation of amylose and the formation of large cyclic alpha-1,4-glucan (cycloamylose) from linear amylose. TLGT functions as a homodimer. Each monomer is composed of two domains, an N-terminal catalytic domain with a (beta/alpha)7 barrel fold and a C-terminal domain with a twisted beta-sandwich fold. Some family members have been designated as alpha-amylases, such as the heat-stable eubacterial amylase from Dictyoglomus thermophilum (DtAmyA) and the extremely thermostable archaeal amylase from Pyrococcus furiosus(PfAmyA). However, both of these proteins are 4-alpha-glucanotransferases. DtAmyA was shown to have transglycosylating activity and PfAmyA exhibits 4-alpha-glucanotransferase activity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD41434.1 | 9.27e-229 | 1 | 431 | 1 | 431 |
| QCP71895.1 | 2.95e-226 | 1 | 431 | 1 | 431 |
| QCD38208.1 | 2.95e-226 | 1 | 431 | 1 | 431 |
| ANU62648.1 | 3.64e-225 | 1 | 440 | 1 | 442 |
| QQR10014.1 | 3.64e-225 | 1 | 440 | 1 | 442 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q59006 | 9.30e-86 | 4 | 440 | 3 | 444 | Putative alpha-amylase OS=Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440) OX=243232 GN=MJ1611 PE=3 SV=1 |
| P09961 | 2.94e-06 | 37 | 170 | 21 | 149 | Alpha-amylase 1 OS=Dictyoglomus thermophilum (strain ATCC 35947 / DSM 3960 / H-6-12) OX=309799 GN=amyA PE=1 SV=2 |
| O57932 | 8.63e-06 | 39 | 163 | 22 | 140 | Alpha-amylase OS=Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) OX=70601 GN=amyA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
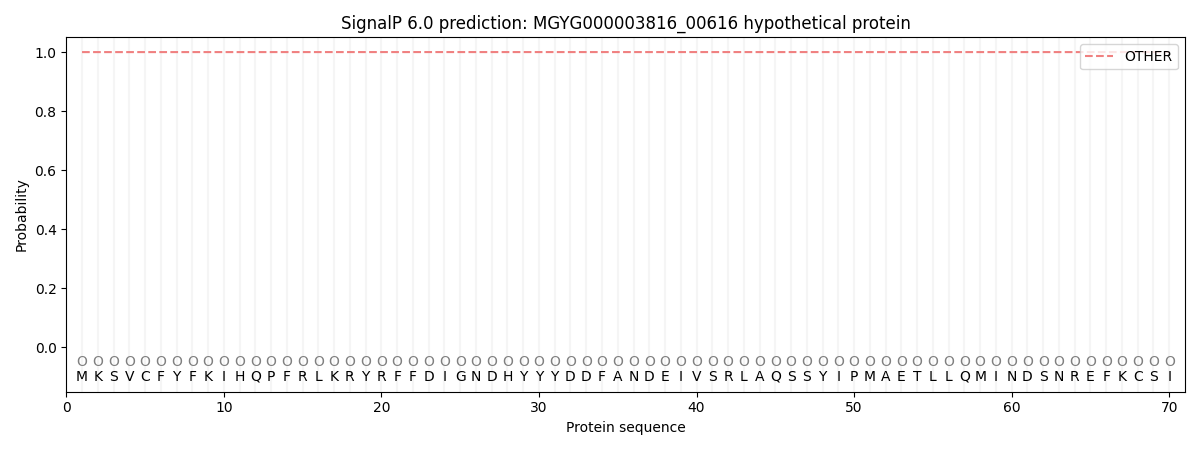
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000063 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
