You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003844_00431
You are here: Home > Sequence: MGYG000003844_00431
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminiclostridium_E sp900539195 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminiclostridium_E; Ruminiclostridium_E sp900539195 | |||||||||||
| CAZyme ID | MGYG000003844_00431 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 172728; End: 175412 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 4 | 403 | 9.8e-53 | 0.46941489361702127 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 3.39e-19 | 5 | 586 | 13 | 583 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 9.66e-07 | 56 | 384 | 59 | 417 | beta-D-glucuronidase; Provisional |
| pfam00703 | Glyco_hydro_2 | 0.001 | 166 | 255 | 2 | 105 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| pfam02836 | Glyco_hydro_2_C | 0.005 | 262 | 384 | 5 | 135 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL34265.1 | 0.0 | 1 | 894 | 1 | 894 |
| CBK95918.1 | 0.0 | 1 | 894 | 1 | 894 |
| BAV13119.1 | 3.32e-257 | 1 | 890 | 1 | 909 |
| ADL53165.1 | 3.32e-257 | 1 | 890 | 1 | 909 |
| CCO04743.1 | 1.65e-254 | 1 | 892 | 1 | 901 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4JKM_A | 3.71e-11 | 5 | 384 | 16 | 415 | CrystalStructure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],4JKM_B Crystal Structure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],6CXS_A Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13],6CXS_B Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13] |
| 6U7I_A | 8.42e-11 | 149 | 384 | 183 | 412 | Faecalibacteriumprausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_B Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_C Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_D Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 6U7J_A | 1.10e-10 | 62 | 390 | 75 | 423 | UnculturedClostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_B Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_C Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_D Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.] |
| 1YQ2_A | 3.91e-09 | 66 | 384 | 122 | 443 | ChainA, beta-galactosidase [Arthrobacter sp. C2-2],1YQ2_B Chain B, beta-galactosidase [Arthrobacter sp. C2-2],1YQ2_C Chain C, beta-galactosidase [Arthrobacter sp. C2-2],1YQ2_D Chain D, beta-galactosidase [Arthrobacter sp. C2-2],1YQ2_E Chain E, beta-galactosidase [Arthrobacter sp. C2-2],1YQ2_F Chain F, beta-galactosidase [Arthrobacter sp. C2-2] |
| 3GM8_A | 7.20e-08 | 4 | 388 | 6 | 400 | ChainA, Glycoside hydrolase family 2, candidate beta-glycosidase [Phocaeicola vulgatus ATCC 8482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KPJ7 | 2.03e-13 | 6 | 386 | 55 | 432 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
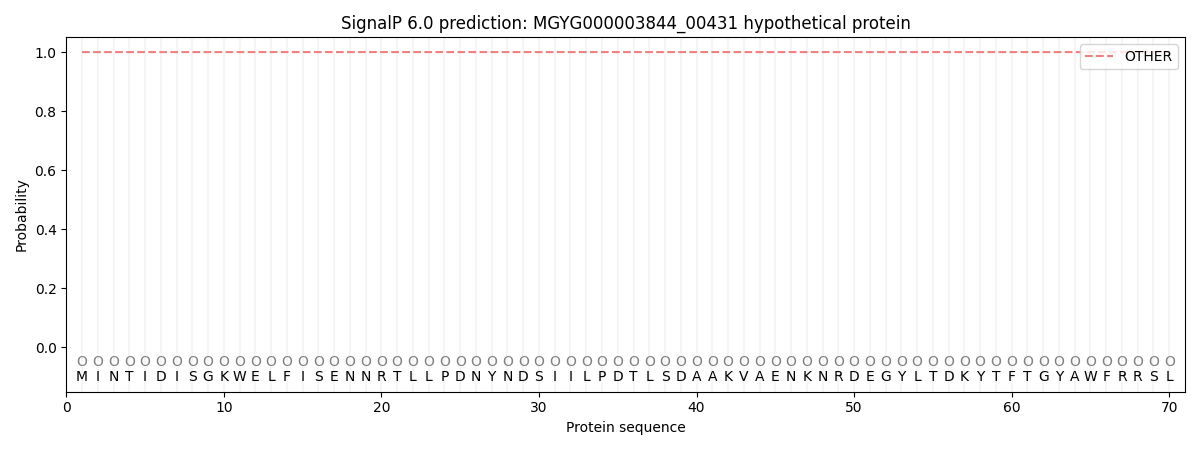
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000056 | 0.000006 | 0.000002 | 0.000000 | 0.000000 | 0.000000 |
