You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003925_02291
You are here: Home > Sequence: MGYG000003925_02291
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
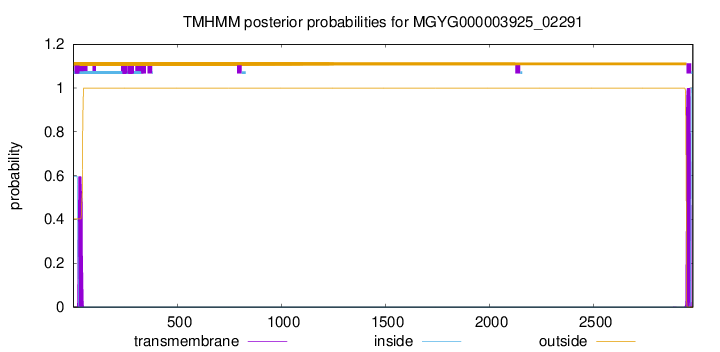
TMHMM annotations
Basic Information help
| Species | CAG-882 sp900545175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; CAG-882; CAG-882 sp900545175 | |||||||||||
| CAZyme ID | MGYG000003925_02291 | |||||||||||
| CAZy Family | CE12 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 840; End: 9776 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL11 | 1018 | 1642 | 7.5e-205 | 0.9588336192109777 |
| CE12 | 250 | 486 | 9.7e-63 | 0.9952380952380953 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10318 | RGL11 | 0.0 | 987 | 1634 | 3 | 564 | Rhamnogalacturonan lyase of the polysaccharide lyase family 11. The rhamnogalacturonan lyase of the polysaccharide lyase family 11 (RGL11) cleaves glycoside bonds in polygalacturonan as well as RG (rhamnogalacturonan) type-I through a beta-elimination reaction. Functionally characterized members of this family, YesW and YesX from Bacillus subtilis, cleave glycoside bonds between rhamnose and galacturonic acid residues in the RG-I region of plant cell wall pectin. YesW and YesX work synergistically, with YesW cleaving the glycoside bond of the RG chain endolytically, and YesX converting the resultant oligosaccharides through an exotype reaction. This domain is sometimes found in architectures with non-catalytic carbohydrate-binding modules (CBMs). There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain through a beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. |
| cd01821 | Rhamnogalacturan_acetylesterase_like | 2.04e-62 | 249 | 486 | 1 | 198 | Rhamnogalacturan_acetylesterase_like subgroup of SGNH-hydrolases. Rhamnogalacturan acetylesterase removes acetyl esters from rhamnogalacturonan substrates, and renders them susceptible to degradation by rhamnogalacturonases. Rhamnogalacturonans are highly branched regions in pectic polysaccharides, consisting of repeating -(1,2)-L-Rha-(1,4)-D-GalUA disaccharide units, with many rhamnose residues substituted by neutral oligosaccharides such as arabinans, galactans and arabinogalactans. Extracellular enzymes participating in the degradation of plant cell wall polymers, such as Rhamnogalacturonan acetylesterase, would typically be found in saprophytic and plant pathogenic fungi and bacteria. |
| NF033839 | PspC_subgroup_2 | 2.15e-17 | 2587 | 2759 | 322 | 512 | pneumococcal surface protein PspC, LPXTG-anchored form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. The other form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. |
| NF033839 | PspC_subgroup_2 | 8.33e-17 | 2587 | 2767 | 338 | 540 | pneumococcal surface protein PspC, LPXTG-anchored form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. The other form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. |
| NF033839 | PspC_subgroup_2 | 2.34e-16 | 2587 | 2761 | 291 | 472 | pneumococcal surface protein PspC, LPXTG-anchored form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. The other form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADL33943.1 | 0.0 | 65 | 2568 | 168 | 2465 |
| BCJ93926.1 | 0.0 | 62 | 2557 | 51 | 2107 |
| QYR24106.1 | 1.00e-281 | 153 | 1642 | 62 | 1388 |
| QNF30995.1 | 1.26e-279 | 110 | 1640 | 46 | 1415 |
| AWB46910.1 | 2.29e-277 | 153 | 1636 | 109 | 1421 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2ZUY_A | 1.05e-154 | 1019 | 1661 | 22 | 614 | Crystalstructure of exotype rhamnogalacturonan lyase YesX [Bacillus subtilis] |
| 2Z8R_A | 2.24e-153 | 1020 | 1640 | 20 | 572 | Crystalstructure of rhamnogalacturonan lyase YesW at 1.40 A resolution [Bacillus subtilis],2Z8R_B Crystal structure of rhamnogalacturonan lyase YesW at 1.40 A resolution [Bacillus subtilis],2Z8S_A Crystal structure of rhamnogalacturonan lyase YesW complexed with digalacturonic acid [Bacillus subtilis],2Z8S_B Crystal structure of rhamnogalacturonan lyase YesW complexed with digalacturonic acid [Bacillus subtilis],2ZUX_A Crystal structure of rhamnogalacturonan lyase YesW complexed with rhamnose [Bacillus subtilis],2ZUX_B Crystal structure of rhamnogalacturonan lyase YesW complexed with rhamnose [Bacillus subtilis] |
| 4CAG_A | 2.71e-147 | 1020 | 1644 | 26 | 583 | Bacilluslicheniformis Rhamnogalacturonan Lyase PL11 [Bacillus licheniformis] |
| 2O14_A | 1.88e-15 | 70 | 447 | 32 | 335 | X-RayCrystal Structure of Protein YXIM_BACsu from Bacillus subtilis. Northeast Structural Genomics Consortium Target SR595 [Bacillus subtilis] |
| 1DEO_A | 9.65e-07 | 250 | 459 | 2 | 201 | RHAMNOGALACTURONANACETYLESTERASE FROM ASPERGILLUS ACULEATUS AT 1.55 A RESOLUTION WITH SO4 IN THE ACTIVE SITE [Aspergillus aculeatus],1DEX_A RHAMNOGALACTURONAN ACETYLESTERASE FROM ASPERGILLUS ACULEATUS AT 1.9 A RESOLUTION [Aspergillus aculeatus],1K7C_A Rhamnogalacturonan acetylesterase with seven N-linked carbohydrate residues distributed at two N-glycosylation sites refined at 1.12 A resolution [Aspergillus aculeatus],1PP4_A The crystal structure of rhamnogalacturonan acetylesterase in space group P3121 [Aspergillus aculeatus],1PP4_B The crystal structure of rhamnogalacturonan acetylesterase in space group P3121 [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O31527 | 8.23e-154 | 1019 | 1640 | 22 | 593 | Rhamnogalacturonan exolyase YesX OS=Bacillus subtilis (strain 168) OX=224308 GN=yesX PE=1 SV=1 |
| O31526 | 3.14e-152 | 1020 | 1640 | 57 | 609 | Rhamnogalacturonan endolyase YesW OS=Bacillus subtilis (strain 168) OX=224308 GN=yesW PE=1 SV=1 |
| O31528 | 1.75e-38 | 251 | 486 | 5 | 202 | Probable rhamnogalacturonan acetylesterase YesY OS=Bacillus subtilis (strain 168) OX=224308 GN=yesY PE=1 SV=1 |
| O31523 | 3.81e-34 | 251 | 486 | 8 | 212 | Rhamnogalacturonan acetylesterase RhgT OS=Bacillus subtilis (strain 168) OX=224308 GN=rhgT PE=1 SV=1 |
| P42304 | 3.40e-16 | 70 | 448 | 47 | 351 | Uncharacterized esterase YxiM OS=Bacillus subtilis (strain 168) OX=224308 GN=yxiM PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
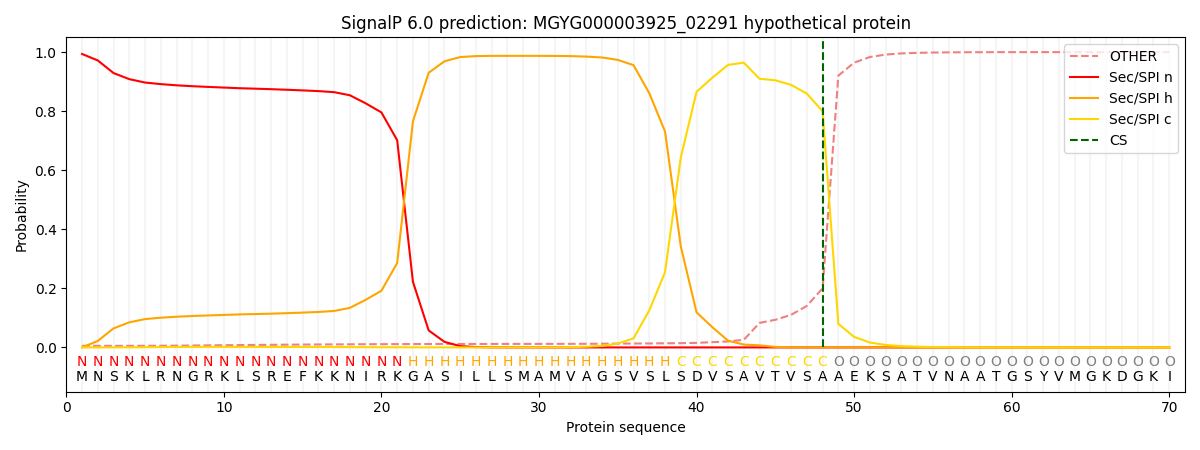
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.006421 | 0.991333 | 0.001184 | 0.000628 | 0.000227 | 0.000181 |