You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004036_01897
You are here: Home > Sequence: MGYG000004036_01897
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1820 sp900545865 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; UMGS1820; UMGS1820 sp900545865 | |||||||||||
| CAZyme ID | MGYG000004036_01897 | |||||||||||
| CAZy Family | GH129 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 67; End: 1947 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH129 | 4 | 624 | 6.1e-131 | 0.9498381877022654 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14244 | GH_101_like | 1.94e-38 | 285 | 519 | 3 | 248 | Endo-a-N-acetylgalactosaminidase and related glcyosyl hydrolases. This family contains the enzymatically active domain of cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins (EC:3.2.1.97). It has been classified as glycosyl hydrolase family 101 in the Cazy resource. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae and other commensal human bacteria is largely determined by their ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. |
| pfam18952 | DUF5696 | 8.62e-10 | 230 | 623 | 209 | 598 | Family of unknown function (DUF5696). This is a family of unknown function with some overlap with clan family members of CL0058. |
| pfam12905 | Glyco_hydro_101 | 1.28e-06 | 314 | 438 | 47 | 178 | Endo-alpha-N-acetylgalactosaminidase. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae is largely determined by the ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. This family is the enzymatic region, EC:3.2.1.97, of the cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins. This reaction is exemplified by the S. pneumoniae protein Endo-alpha-N-acetylgalactosaminidase, where Asp764 is the catalytic nucleophile-base and Glu796 the catalytic proton donor. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUH27706.1 | 3.92e-179 | 3 | 623 | 7 | 621 |
| QUH27660.1 | 4.28e-163 | 46 | 623 | 56 | 626 |
| QNK55038.1 | 2.75e-146 | 66 | 625 | 81 | 626 |
| QUO33311.1 | 1.39e-143 | 14 | 623 | 14 | 627 |
| QJD87097.1 | 1.75e-140 | 74 | 625 | 92 | 627 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5WZN_A | 8.08e-121 | 39 | 624 | 40 | 629 | Alpha-N-acetylgalactosaminidaseNagBb from Bifidobacterium bifidum - GalNAc complex [Bifidobacterium bifidum],5WZN_B Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - GalNAc complex [Bifidobacterium bifidum],5WZP_A Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - ligand free [Bifidobacterium bifidum],5WZP_B Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - ligand free [Bifidobacterium bifidum],5WZR_A Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - Gal-NHAc-DNJ complex [Bifidobacterium bifidum],5WZR_B Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - Gal-NHAc-DNJ complex [Bifidobacterium bifidum] |
| 5WZQ_A | 3.58e-115 | 39 | 624 | 40 | 629 | Alpha-N-acetylgalactosaminidaseNagBb from Bifidobacterium bifidum - quadruple mutant [Bifidobacterium bifidum],5WZQ_B Alpha-N-acetylgalactosaminidase NagBb from Bifidobacterium bifidum - quadruple mutant [Bifidobacterium bifidum] |
| 5OPQ_A | 1.08e-15 | 230 | 623 | 299 | 661 | A3,6-anhydro-D-galactosidase produced by Zobellia galactanivorans. This is an exo-lytic enzyme that hydrolyzes terminal 3,6-anhydro-D-galactose from the non-reducing end of carrageenan oligosaccharides. [Zobellia galactanivorans],5OPQ_B A 3,6-anhydro-D-galactosidase produced by Zobellia galactanivorans. This is an exo-lytic enzyme that hydrolyzes terminal 3,6-anhydro-D-galactose from the non-reducing end of carrageenan oligosaccharides. [Zobellia galactanivorans],5OPQ_C A 3,6-anhydro-D-galactosidase produced by Zobellia galactanivorans. This is an exo-lytic enzyme that hydrolyzes terminal 3,6-anhydro-D-galactose from the non-reducing end of carrageenan oligosaccharides. [Zobellia galactanivorans],5OPQ_D A 3,6-anhydro-D-galactosidase produced by Zobellia galactanivorans. This is an exo-lytic enzyme that hydrolyzes terminal 3,6-anhydro-D-galactose from the non-reducing end of carrageenan oligosaccharides. [Zobellia galactanivorans] |
| 6QEP_A | 3.38e-07 | 299 | 438 | 317 | 459 | EngBFDARPin Fusion 4b H14 [Bifidobacterium longum] |
| 6QEV_B | 3.38e-07 | 299 | 438 | 317 | 459 | EngBFDARPin Fusion 4b B6 [Bifidobacterium longum] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
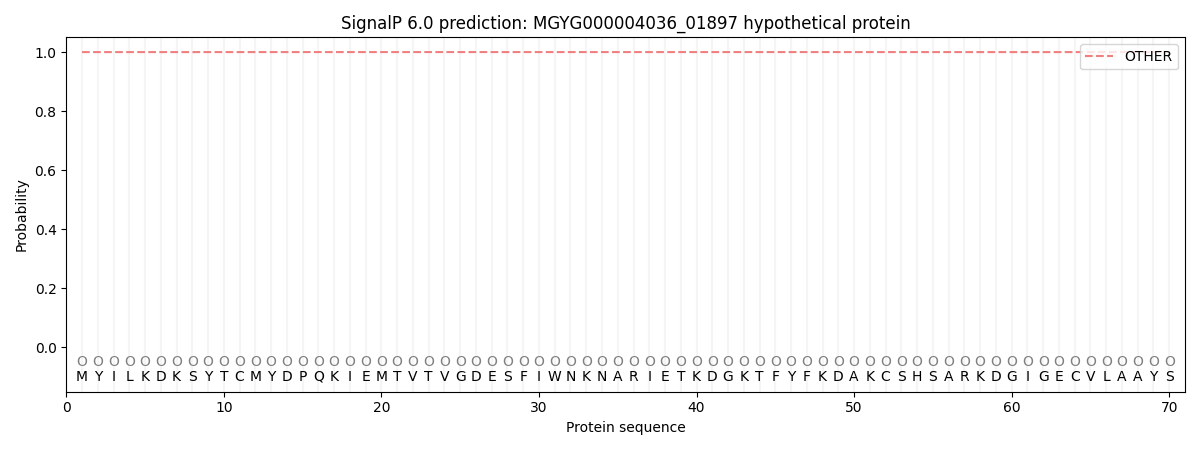
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000081 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
