You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004053_01763
You are here: Home > Sequence: MGYG000004053_01763
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; | |||||||||||
| CAZyme ID | MGYG000004053_01763 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Exo-beta-D-glucosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1222; End: 3501 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 26 | 710 | 1.2e-78 | 0.6728723404255319 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 4.79e-45 | 13 | 488 | 21 | 445 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10340 | ebgA | 1.43e-12 | 213 | 451 | 220 | 449 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam00703 | Glyco_hydro_2 | 8.25e-11 | 215 | 309 | 13 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK09525 | lacZ | 1.98e-07 | 213 | 451 | 232 | 462 | beta-galactosidase. |
| pfam02836 | Glyco_hydro_2_C | 3.34e-06 | 336 | 451 | 7 | 135 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI62071.1 | 3.87e-257 | 1 | 754 | 1 | 744 |
| BBL06539.1 | 1.56e-246 | 23 | 755 | 9 | 728 |
| AUX47279.1 | 3.74e-211 | 15 | 752 | 12 | 727 |
| QUI21034.1 | 3.61e-210 | 24 | 755 | 6 | 727 |
| ACB77359.1 | 1.88e-175 | 23 | 750 | 15 | 741 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5N6U_A | 1.37e-58 | 44 | 652 | 46 | 656 | Crystalstructure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_B Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_C Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_D Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
| 2VJX_A | 1.77e-55 | 40 | 759 | 24 | 759 | Structuraland biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VJX_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VQT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VQT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VR4_A Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482],2VR4_B Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482] |
| 2JE8_A | 1.80e-55 | 40 | 759 | 26 | 761 | Structureof a beta-mannosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],2JE8_B Structure of a beta-mannosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
| 7OP6_A | 1.81e-55 | 40 | 759 | 26 | 761 | ChainA, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP6_B Chain B, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP7_A Chain A, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP7_B Chain B, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482] |
| 2WBK_A | 4.45e-55 | 40 | 759 | 24 | 759 | Structureof the Michaelis complex of beta-mannosidase, Man2A, provides insight into the conformational itinerary of mannoside hydrolysis [Bacteroides thetaiotaomicron VPI-5482],2WBK_B Structure of the Michaelis complex of beta-mannosidase, Man2A, provides insight into the conformational itinerary of mannoside hydrolysis [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| I2C092 | 8.81e-54 | 25 | 652 | 12 | 660 | Beta-mannosidase B OS=Thermothelomyces thermophilus OX=78579 GN=man9 PE=1 SV=1 |
| B8NW36 | 3.44e-51 | 23 | 663 | 8 | 667 | Beta-mannosidase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=mndB PE=3 SV=1 |
| Q2TXB7 | 4.65e-51 | 23 | 663 | 8 | 667 | Beta-mannosidase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=mndB PE=3 SV=3 |
| A1CGA8 | 6.21e-48 | 29 | 663 | 12 | 667 | Beta-mannosidase B OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=mndB PE=3 SV=1 |
| Q5B7W2 | 7.47e-45 | 45 | 663 | 31 | 667 | Beta-mannosidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=mndB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
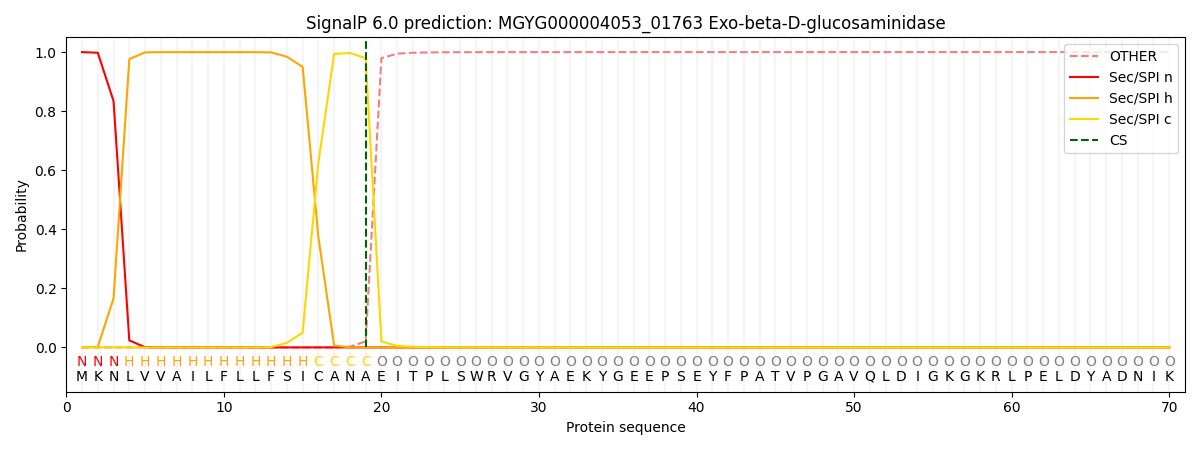
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000248 | 0.999042 | 0.000211 | 0.000160 | 0.000151 | 0.000142 |
