You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004128_01199
You are here: Home > Sequence: MGYG000004128_01199
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1253 sp900550065 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UMGS1253; UMGS1253; UMGS1253 sp900550065 | |||||||||||
| CAZyme ID | MGYG000004128_01199 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 767; End: 2029 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 48 | 274 | 3.4e-32 | 0.9869565217391304 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06438 | EpsO_like | 2.95e-68 | 51 | 233 | 1 | 183 | EpsO protein participates in the methanolan synthesis. The Methylobacillus sp EpsO protein is predicted to participate in the methanolan synthesis. Methanolan is an exopolysaccharide (EPS), composed of glucose, mannose and galactose. A 21 genes cluster was predicted to participate in the methanolan synthesis. Gene disruption analysis revealed that EpsO is one of the glycosyltransferase enzymes involved in the synthesis of repeating sugar units onto the lipid carrier. |
| COG1215 | BcsA | 8.89e-38 | 1 | 397 | 1 | 406 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| cd06423 | CESA_like | 1.81e-33 | 51 | 230 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| pfam13641 | Glyco_tranf_2_3 | 2.53e-26 | 48 | 271 | 3 | 227 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| cd04179 | DPM_DPG-synthase_like | 2.30e-17 | 51 | 219 | 1 | 165 | DPM_DPG-synthase_like is a member of the Glycosyltransferase 2 superfamily. DPM1 is the catalytic subunit of eukaryotic dolichol-phosphate mannose (DPM) synthase. DPM synthase is required for synthesis of the glycosylphosphatidylinositol (GPI) anchor, N-glycan precursor, protein O-mannose, and C-mannose. In higher eukaryotes,the enzyme has three subunits, DPM1, DPM2 and DPM3. DPM is synthesized from dolichol phosphate and GDP-Man on the cytosolic surface of the ER membrane by DPM synthase and then is flipped onto the luminal side and used as a donor substrate. In lower eukaryotes, such as Saccharomyces cerevisiae and Trypanosoma brucei, DPM synthase consists of a single component (Dpm1p and TbDpm1, respectively) that possesses one predicted transmembrane region near the C terminus for anchoring to the ER membrane. In contrast, the Dpm1 homologues of higher eukaryotes, namely fission yeast, fungi, and animals, have no transmembrane region, suggesting the existence of adapter molecules for membrane anchoring. This family also includes bacteria and archaea DPM1_like enzymes. However, the enzyme structure and mechanism of function are not well understood. The UDP-glucose:dolichyl-phosphate glucosyltransferase (DPG_synthase) is a transmembrane-bound enzyme of the endoplasmic reticulum involved in protein N-linked glycosylation. This enzyme catalyzes the transfer of glucose from UDP-glucose to dolichyl phosphate. This protein family belongs to Glycosyltransferase 2 superfamily. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALU13522.1 | 2.37e-118 | 8 | 417 | 5 | 410 |
| QCT71577.1 | 2.37e-118 | 8 | 417 | 5 | 410 |
| ADO38773.1 | 4.74e-118 | 8 | 417 | 5 | 410 |
| ARD65366.1 | 4.74e-118 | 8 | 417 | 5 | 410 |
| AWW28570.1 | 7.16e-113 | 52 | 418 | 1 | 363 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96587 | 4.62e-10 | 35 | 274 | 36 | 279 | Uncharacterized glycosyltransferase YdaM OS=Bacillus subtilis (strain 168) OX=224308 GN=ydaM PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
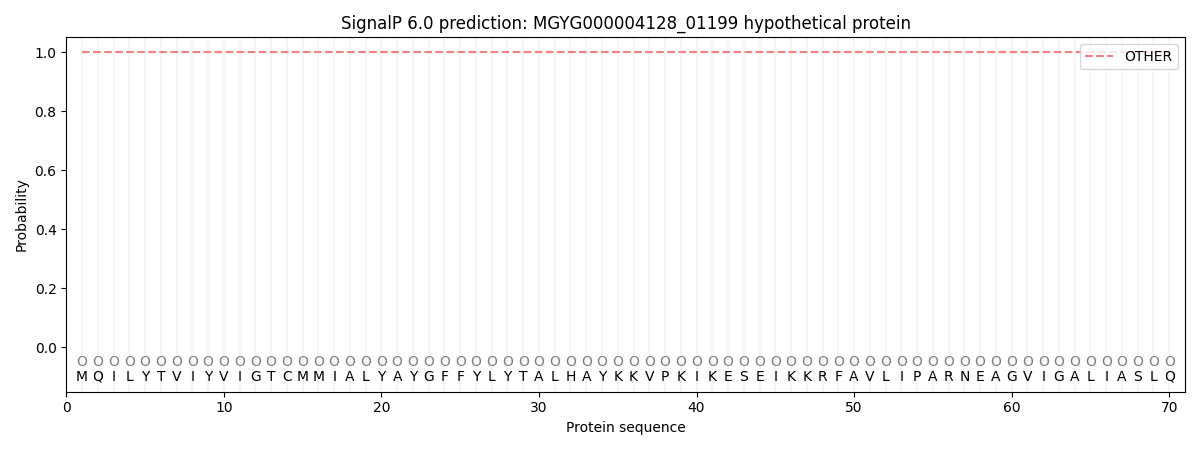
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000038 | 0.000003 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |

