You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004128_01724
You are here: Home > Sequence: MGYG000004128_01724
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1253 sp900550065 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UMGS1253; UMGS1253; UMGS1253 sp900550065 | |||||||||||
| CAZyme ID | MGYG000004128_01724 | |||||||||||
| CAZy Family | PL21 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1288; End: 4071 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL21 | 734 | 798 | 3.3e-24 | 0.9444444444444444 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd04086 | CBM35_mannanase-like | 1.88e-05 | 202 | 308 | 3 | 119 | Carbohydrate Binding Module 35 (CBM35); appended to several carbohydrate binding enzymes, including several glycoside hydrolase (GH) family 26 mannanase domains. This family includes carbohydrate binding module 35 (CBM35) domains that are appended to several carbohydrate binding enzymes, including periplasmic component of ABC-type sugar transport system involved in carbohydrate transport and metabolism, and several glycoside hydrolase (GH) domains, including GH26. These CBM6s are non-catalytic carbohydrate binding domains that facilitate the strong binding of the GH catalytic modules with their dedicated, insoluble substrates. Examples of proteins having CMB35s belonging to this family are mannanase A from Clostridium thermocellum (GH26), Man26B from Paenibacillus sp. BME-14 (GH26), and the multifunctional Cel44C-Man26A from Paenibacillus polymyxa GS01 (which has two GH domains, GH44 and GH26). GH26 mainly includes mannan endo-1,4-beta-mannosidase which hydrolyzes 1,4-beta-D-linkages in mannans, galacto-mannans, glucomannans, and galactoglucomannans, but displays little activity towards other plant cell wall polysaccharides. A few proteins belonging to this family have additional CBM3 domains; these CBM3s are not found in the CBM6-CBM35-CBM36_like superfamily. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI63941.1 | 2.28e-164 | 362 | 927 | 32 | 614 |
| BCI63940.1 | 7.00e-154 | 367 | 927 | 41 | 617 |
| BBH20141.1 | 3.49e-136 | 362 | 927 | 228 | 787 |
| QHW00380.1 | 1.15e-122 | 368 | 927 | 66 | 632 |
| QDG70382.1 | 3.70e-121 | 227 | 926 | 80 | 783 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2FUQ_A | 7.07e-119 | 368 | 927 | 18 | 577 | ChainA, heparinase II protein [Pedobacter heparinus],2FUQ_B Chain B, heparinase II protein [Pedobacter heparinus] |
| 3E80_A | 7.45e-119 | 368 | 927 | 20 | 579 | Structureof Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus],3E80_B Structure of Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus],3E80_C Structure of Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus] |
| 3E7J_A | 4.37e-116 | 368 | 927 | 20 | 579 | ChainA, Heparinase II protein [Pedobacter heparinus],3E7J_B Chain B, Heparinase II protein [Pedobacter heparinus] |
| 2FUT_A | 1.26e-113 | 368 | 927 | 19 | 578 | ChainA, heparinase II protein [Pedobacter heparinus],2FUT_B Chain B, heparinase II protein [Pedobacter heparinus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| C6XZB6 | 5.29e-118 | 329 | 927 | 11 | 602 | Heparin and heparin-sulfate lyase OS=Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / CIP 104194 / JCM 7457 / NBRC 12017 / NCIMB 9290 / NRRL B-14731 / HIM 762-3) OX=485917 GN=hepB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
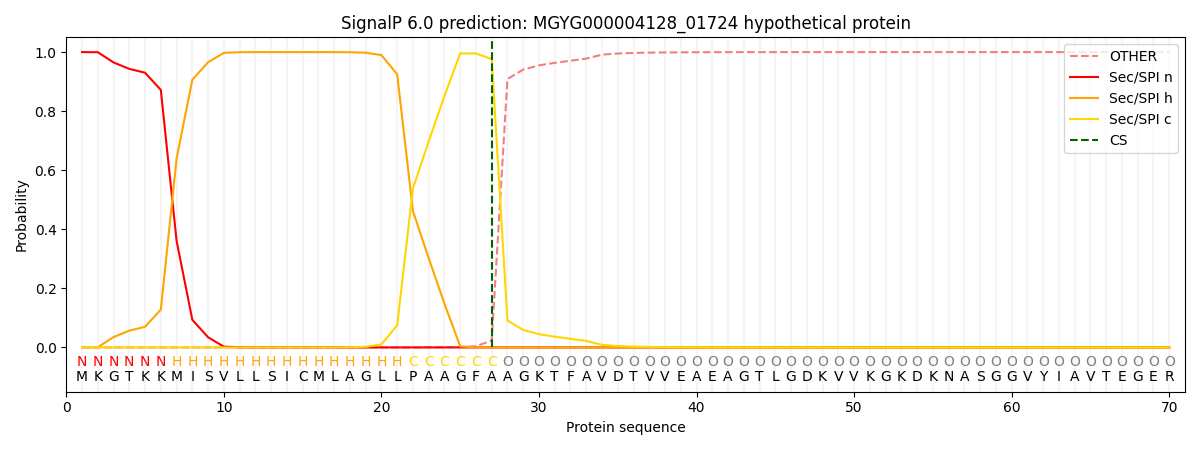
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000263 | 0.998965 | 0.000209 | 0.000195 | 0.000176 | 0.000160 |