You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004174_01331
You are here: Home > Sequence: MGYG000004174_01331
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
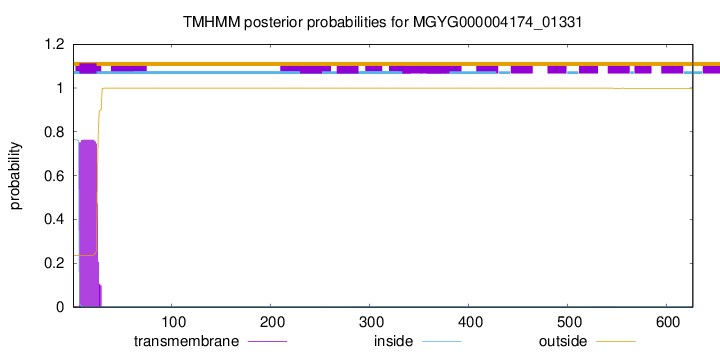
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; Firm-11; | |||||||||||
| CAZyme ID | MGYG000004174_01331 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-glucosidase BoGH3B | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 753; End: 2636 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 134 | 350 | 3.1e-53 | 0.9722222222222222 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 2.93e-49 | 115 | 454 | 35 | 373 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| PRK15098 | PRK15098 | 1.43e-46 | 136 | 625 | 100 | 651 | beta-glucosidase BglX. |
| pfam00933 | Glyco_hydro_3 | 1.74e-42 | 99 | 379 | 25 | 314 | Glycosyl hydrolase family 3 N terminal domain. |
| pfam01915 | Glyco_hydro_3_C | 6.58e-16 | 437 | 621 | 22 | 215 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| PLN03080 | PLN03080 | 4.73e-12 | 197 | 625 | 155 | 636 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QTE67436.1 | 1.25e-315 | 1 | 627 | 1 | 657 |
| QTE67433.1 | 1.27e-127 | 25 | 620 | 29 | 656 |
| AXH35175.1 | 7.00e-83 | 31 | 620 | 39 | 650 |
| ARJ06162.1 | 9.77e-83 | 31 | 620 | 39 | 650 |
| QHM76280.1 | 5.69e-79 | 26 | 620 | 31 | 677 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5M6G_A | 2.79e-51 | 60 | 620 | 39 | 612 | Crystalstructure Glucan 1,4-beta-glucosidase from Saccharopolyspora erythraea [Saccharopolyspora erythraea D] |
| 6JGL_A | 9.09e-51 | 61 | 627 | 9 | 608 | Crystalstructure of barley exohydrolaseI W434H mutant in complex with methyl 2-thio-beta-sophoroside [Hordeum vulgare subsp. vulgare],6JGN_A Crystal structure of barley exohydrolaseI W434H in complex with 4'-nitrophenyl thiolaminaribioside [Hordeum vulgare subsp. vulgare],6JGO_A Crystal structure of barley exohydrolaseI W434H mutant in complex with 4I,4III,4V-S-trithiocellohexaose [Hordeum vulgare subsp. vulgare],6JGP_A Crystal structure of barley exohydrolaseI W434H mutant in complex with methyl 6-thio-beta-gentiobioside. [Hordeum vulgare subsp. vulgare] |
| 6JGE_A | 1.25e-50 | 61 | 627 | 9 | 608 | Crystalstructure of barley exohydrolaseI W434A mutant in complex with methyl 2-thio-beta-sophoroside. [Hordeum vulgare subsp. vulgare],6K6V_A Crystal structure of barley exohydrolaseI W434A mutant in complex with methyl 6-thio-beta-gentiobioside [Hordeum vulgare subsp. vulgare],6KUF_A Crystal structure of barley exohydrolaseI W434A mutant in complex with glucose. [Hordeum vulgare subsp. vulgare],6L1J_A Crystal structure of barley exohydrolaseI W434A mutant in complex with 4'-nitrophenyl thiolaminaritrioside [Hordeum vulgare subsp. vulgare],6LBB_A Crystal structure of barley exohydrolaseI W434A mutant in complex with 4I,4III,4V-S-trithiocellohexaose [Hordeum vulgare subsp. vulgare] |
| 6JGQ_A | 1.72e-50 | 61 | 627 | 9 | 608 | Crystalstructure of barley exohydrolaseI W434Y mutant in complex with methyl 2-thio-beta-sophoroside. [Hordeum vulgare subsp. vulgare],6JGR_A Crystal structure of barley exohydrolaseI W434Y in complex with 4'-nitrophenyl thiolaminaribioside [Hordeum vulgare subsp. vulgare],6JGS_A Crystal structure of barley exohydrolaseI W434Y mutant in complex with 4I,4III,4V-S-trithiocellohexaose. [Hordeum vulgare subsp. vulgare],6JGT_A Crystal structure of barley exohydrolaseI W434Y mutant in complex with methyl 6-thio-beta-gentiobioside. [Hordeum vulgare subsp. vulgare] |
| 1EX1_A | 2.22e-50 | 61 | 627 | 5 | 604 | Beta-d-glucanExohydrolase From Barley [Hordeum vulgare],1IEQ_A Crystal Structure Of Barley Beta-D-Glucan Glucohydrolase Isoenzyme Exo1 [Hordeum vulgare],1IEV_A Crystal Structure Of Barley Beta-d-glucan Glucohydrolase Isoenzyme Exo1 In Complex With Cyclohexitol [Hordeum vulgare],1IEW_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 2-deoxy-2-fluoro-alpha-D-glucoside [Hordeum vulgare],1IEX_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 4I,4III,4V-S-trithiocellohexaose [Hordeum vulgare],1J8V_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 4'-nitrophenyl 3I-thiolaminaritrioside [Hordeum vulgare],3WLH_A Crystal Structure Analysis of Plant Exohydrolase [Hordeum vulgare subsp. vulgare],3WLJ_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with 3-deoxy-glucose [Hordeum vulgare subsp. vulgare],3WLK_X Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with 4-deoxy-glucose [Hordeum vulgare subsp. vulgare],3WLL_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with PEG400 [Hordeum vulgare subsp. vulgare],3WLM_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme exo1 in complex with octyl-O-glucoside [Hordeum vulgare subsp. vulgare],3WLN_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with octyl-S-glucoside [Hordeum vulgare subsp. vulgare],3WLP_A Crystal Structure Analysis of Plant Exohydrolase [Hordeum vulgare subsp. vulgare] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q46684 | 1.56e-70 | 30 | 620 | 27 | 652 | Periplasmic beta-glucosidase/beta-xylosidase OS=Dickeya chrysanthemi OX=556 GN=bgxA PE=3 SV=1 |
| Q2UFP8 | 2.65e-59 | 46 | 620 | 33 | 624 | Probable beta-glucosidase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bglC PE=3 SV=2 |
| B8NGU6 | 6.54e-59 | 61 | 620 | 42 | 620 | Probable beta-glucosidase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglC PE=3 SV=1 |
| Q5BCC6 | 6.83e-59 | 58 | 621 | 35 | 614 | Beta-glucosidase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglC PE=1 SV=1 |
| A7LXU3 | 4.87e-39 | 123 | 625 | 108 | 662 | Beta-glucosidase BoGH3B OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02659 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
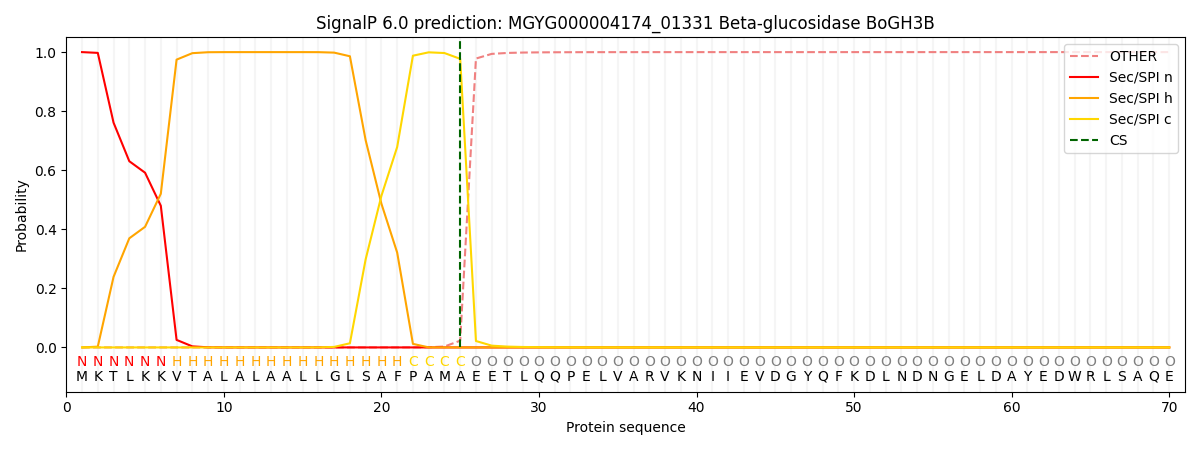
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000236 | 0.999038 | 0.000188 | 0.000183 | 0.000171 | 0.000146 |