You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004336_00692
You are here: Home > Sequence: MGYG000004336_00692
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Faecalibacterium sp900765105 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Faecalibacterium; Faecalibacterium sp900765105 | |||||||||||
| CAZyme ID | MGYG000004336_00692 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-glucuronidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4866; End: 6662 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 11 | 594 | 7.7e-107 | 0.6143617021276596 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10150 | PRK10150 | 0.0 | 6 | 595 | 1 | 592 | beta-D-glucuronidase; Provisional |
| COG3250 | LacZ | 5.00e-105 | 8 | 596 | 1 | 596 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| pfam02836 | Glyco_hydro_2_C | 8.12e-78 | 279 | 596 | 1 | 298 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK10340 | ebgA | 1.91e-40 | 73 | 442 | 113 | 468 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK09525 | lacZ | 2.70e-38 | 16 | 418 | 51 | 462 | beta-galactosidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCZ68610.1 | 0.0 | 1 | 596 | 1 | 597 |
| QRT31041.1 | 0.0 | 1 | 596 | 1 | 597 |
| AAQ76046.1 | 0.0 | 1 | 596 | 1 | 597 |
| QBE95891.1 | 1.11e-279 | 1 | 598 | 1 | 599 |
| QIB58662.1 | 2.49e-277 | 6 | 598 | 1 | 594 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6EC6_A | 0.0 | 1 | 596 | 24 | 620 | Ruminococcusgnavus Beta-glucuronidase [[Ruminococcus] gnavus],6EC6_B Ruminococcus gnavus Beta-glucuronidase [[Ruminococcus] gnavus] |
| 6JZ1_A | 0.0 | 1 | 596 | 1 | 597 | Apostructure of b-glucuronidase from Ruminococcus gnavus at 1.7 Angstrom resolution [[Ruminococcus] gnavus],6JZ1_B Apo structure of b-glucuronidase from Ruminococcus gnavus at 1.7 Angstrom resolution [[Ruminococcus] gnavus],6JZ5_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucuronic acid [[Ruminococcus] gnavus],6JZ5_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucuronic acid [[Ruminococcus] gnavus],6JZ6_A b-glucuronidase from Ruminococcus gnavus in complex with C6-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ6_B b-glucuronidase from Ruminococcus gnavus in complex with C6-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ7_A b-glucuronidase from Ruminococcus gnavus in complex with N1-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ7_B b-glucuronidase from Ruminococcus gnavus in complex with N1-substituted uronic isofagomine [[Ruminococcus] gnavus],6JZ8_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro 1,5-lactone [[Ruminococcus] gnavus],6JZ8_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro 1,5-lactone [[Ruminococcus] gnavus] |
| 5Z18_A | 0.0 | 1 | 596 | 25 | 621 | Thecrystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_B The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_C The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_D The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_E The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z18_F The crystal structure of Ruminococcus gnavus beta-glucuronidase [[Ruminococcus] gnavus],5Z19_A The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_B The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_C The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_D The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_E The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],5Z19_F The crystal structure of Ruminococcus gnavus beta-glucuronidase in complex with uronic isofagomine [[Ruminococcus] gnavus],6JZ2_A b-glucuronidase from Ruminococcus gnavus in complex with uronic isofagomine at 1.3 Angstroms resolution [[Ruminococcus] gnavus],6JZ2_B b-glucuronidase from Ruminococcus gnavus in complex with uronic isofagomine at 1.3 Angstroms resolution [[Ruminococcus] gnavus],6JZ3_A b-glucuronidase from Ruminococcus gnavus in complex with uronic deoxynojirimycin [[Ruminococcus] gnavus],6JZ3_B b-glucuronidase from Ruminococcus gnavus in complex with uronic deoxynojirimycin [[Ruminococcus] gnavus],6JZ4_A b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro-d-lactam [[Ruminococcus] gnavus],6JZ4_B b-glucuronidase from Ruminococcus gnavus in complex with D-glucaro-d-lactam [[Ruminococcus] gnavus] |
| 6ED2_A | 1.57e-260 | 2 | 598 | 24 | 622 | ChainA, Glycosyl hydrolase family 2, TIM barrel domain protein [Faecalibacterium duncaniae] |
| 7KGY_A | 7.50e-258 | 1 | 598 | 1 | 600 | ChainA, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_B Chain B, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_C Chain C, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_D Chain D, Beta-glucuronidase [Faecalibacterium prausnitzii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P05804 | 1.64e-163 | 6 | 595 | 1 | 588 | Beta-glucuronidase OS=Escherichia coli (strain K12) OX=83333 GN=uidA PE=1 SV=2 |
| O77695 | 4.17e-128 | 2 | 595 | 20 | 623 | Beta-glucuronidase (Fragment) OS=Chlorocebus aethiops OX=9534 GN=GUSB PE=2 SV=1 |
| O97524 | 2.54e-127 | 2 | 595 | 23 | 625 | Beta-glucuronidase OS=Felis catus OX=9685 GN=GUSB PE=1 SV=1 |
| P08236 | 7.11e-127 | 2 | 595 | 23 | 626 | Beta-glucuronidase OS=Homo sapiens OX=9606 GN=GUSB PE=1 SV=2 |
| O18835 | 1.00e-126 | 2 | 595 | 23 | 625 | Beta-glucuronidase OS=Canis lupus familiaris OX=9615 GN=GUSB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
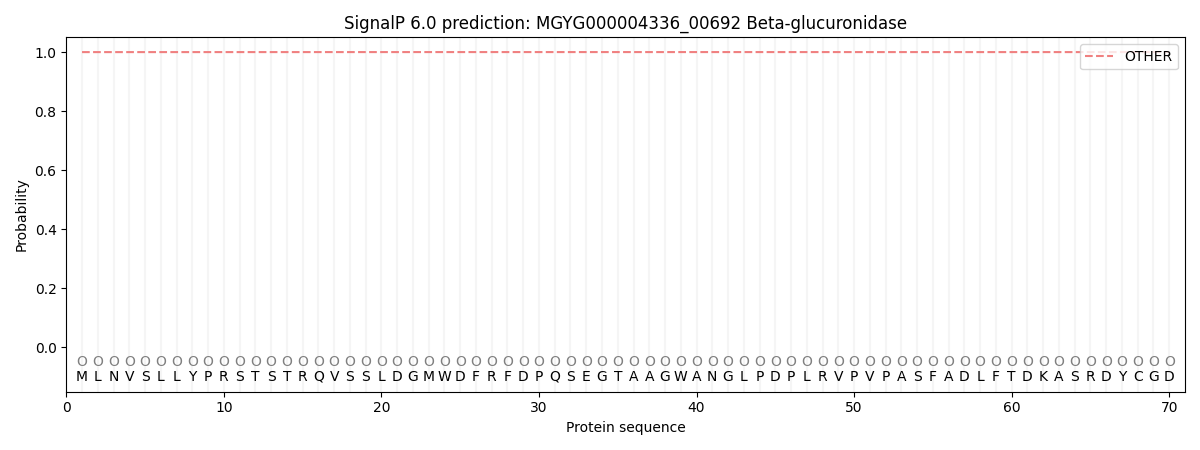
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000048 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
