You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004384_00657
You are here: Home > Sequence: MGYG000004384_00657
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
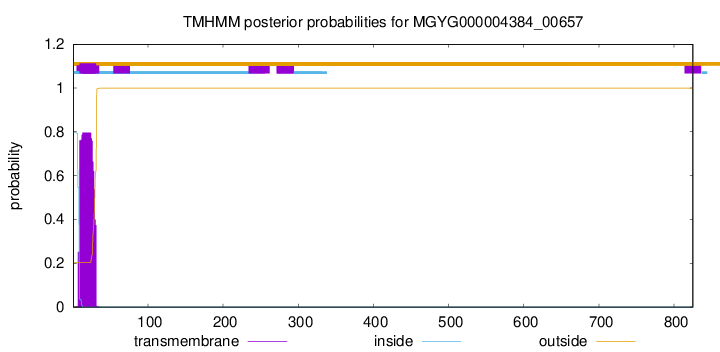
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminococcus; | |||||||||||
| CAZyme ID | MGYG000004384_00657 | |||||||||||
| CAZy Family | GH48 | |||||||||||
| CAZyme Description | Cellulose 1,4-beta-cellobiosidase (reducing end) CelS | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 856; End: 3333 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH48 | 45 | 725 | 2.1e-206 | 0.9950900163666121 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02011 | Glyco_hydro_48 | 0.0 | 45 | 728 | 1 | 620 | Glycosyl hydrolase family 48. Members of this family are endoglucanase EC:3.2.1.4 and exoglucanase EC:3.2.1.91 enzymes that cleave cellulose or related substrate. |
| cd14256 | Dockerin_I | 2.41e-05 | 744 | 811 | 2 | 57 | Type I dockerin repeat domain. Bacterial cohesin domains bind to a complementary protein domain named dockerin, and this interaction is required for the formation of the cellulosome, a cellulose-degrading complex. The cellulosome consists of scaffoldin, a noncatalytic scaffolding polypeptide, that comprises repeating cohesion modules and a single carbohydrate-binding module (CBM). Specific calcium-dependent interactions between cohesins and dockerins appear to be essential for cellulosome assembly. This subfamily represents type I dockerins, which are responsible for anchoring a variety of enzymatic domains to the complex. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADX05724.1 | 0.0 | 1 | 767 | 1 | 806 |
| CBL17316.1 | 2.15e-290 | 1 | 815 | 1 | 798 |
| ADU23081.1 | 2.97e-250 | 1 | 772 | 1 | 815 |
| BAJ05814.1 | 2.97e-250 | 1 | 772 | 1 | 815 |
| CAS03459.1 | 7.49e-250 | 1 | 772 | 1 | 821 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6D5D_A | 5.08e-165 | 63 | 728 | 36 | 642 | Structureof Caldicellulosiruptor danielii GH48 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii] |
| 1L1Y_A | 2.02e-164 | 13 | 731 | 10 | 664 | ChainA, cellobiohydrolase [Acetivibrio thermocellus],1L1Y_B Chain B, cellobiohydrolase [Acetivibrio thermocellus],1L1Y_C Chain C, cellobiohydrolase [Acetivibrio thermocellus],1L1Y_D Chain D, cellobiohydrolase [Acetivibrio thermocellus],1L1Y_E Chain E, cellobiohydrolase [Acetivibrio thermocellus],1L1Y_F Chain F, cellobiohydrolase [Acetivibrio thermocellus],1L2A_A Chain A, cellobiohydrolase [Acetivibrio thermocellus],1L2A_B Chain B, cellobiohydrolase [Acetivibrio thermocellus],1L2A_C Chain C, cellobiohydrolase [Acetivibrio thermocellus],1L2A_D Chain D, cellobiohydrolase [Acetivibrio thermocellus],1L2A_E Chain E, cellobiohydrolase [Acetivibrio thermocellus],1L2A_F Chain F, cellobiohydrolase [Acetivibrio thermocellus] |
| 5YJ6_A | 7.76e-164 | 44 | 730 | 12 | 636 | ChainA, Dockerin type I repeat-containing protein [Acetivibrio thermocellus DSM 1313] |
| 4EL8_A | 9.86e-162 | 63 | 728 | 25 | 631 | Theunliganded structure of C.bescii CelA GH48 module [Caldicellulosiruptor bescii DSM 6725] |
| 4L0G_A | 1.97e-161 | 63 | 728 | 29 | 635 | CrystalStructure of a GH48 cellobiohydrolase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii DSM 6725],4L6X_A Crystal Structure of a GH48 cellobiohydrolase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii DSM 6725],4TXT_A Crystal Structure of a GH48 cellobiohydrolase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DH67 | 6.44e-168 | 13 | 760 | 10 | 694 | Cellulose 1,4-beta-cellobiosidase (reducing end) CelS OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celS PE=1 SV=1 |
| P0C2S5 | 6.44e-168 | 13 | 760 | 10 | 694 | Cellulose 1,4-beta-cellobiosidase (reducing end) CelS OS=Acetivibrio thermocellus OX=1515 GN=celS PE=1 SV=1 |
| P37698 | 5.48e-164 | 2 | 766 | 1 | 688 | Endoglucanase F OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCF PE=1 SV=2 |
| P50900 | 6.65e-155 | 42 | 728 | 37 | 655 | Exoglucanase-2 OS=Thermoclostridium stercorarium (strain ATCC 35414 / DSM 8532 / NCIMB 11754) OX=1121335 GN=celY PE=1 SV=2 |
| P22534 | 4.81e-153 | 76 | 728 | 1140 | 1739 | Endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
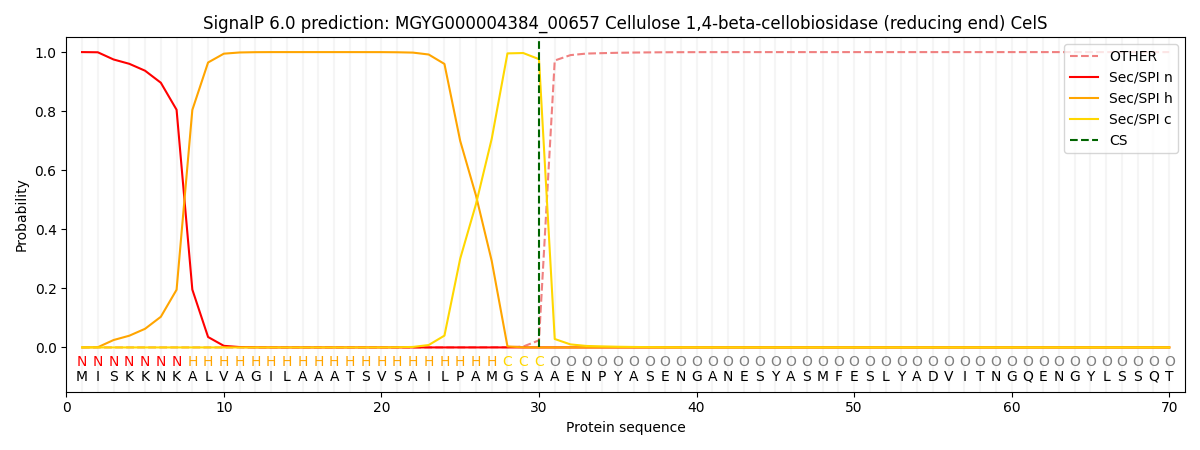
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000218 | 0.999080 | 0.000174 | 0.000191 | 0.000167 | 0.000137 |