You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004400_00093
You are here: Home > Sequence: MGYG000004400_00093
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; RF32; CAG-239; CAG-495; | |||||||||||
| CAZyme ID | MGYG000004400_00093 | |||||||||||
| CAZy Family | GT30 | |||||||||||
| CAZyme Description | 3-deoxy-D-manno-octulosonic acid transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 94115; End: 95374 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT30 | 34 | 209 | 1.1e-62 | 0.9774011299435028 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05749 | PRK05749 | 1.27e-144 | 1 | 419 | 1 | 421 | 3-deoxy-D-manno-octulosonic-acid transferase; Reviewed |
| COG1519 | KdtA | 1.43e-139 | 2 | 419 | 1 | 419 | 3-deoxy-D-manno-octulosonic-acid transferase [Cell wall/membrane/envelope biogenesis]. |
| pfam04413 | Glycos_transf_N | 1.55e-75 | 38 | 210 | 2 | 175 | 3-Deoxy-D-manno-octulosonic-acid transferase (kdotransferase). Members of this family transfer activated sugars to a variety of substrates, including glycogen, fructose-6-phosphate and lipopolysaccharides. Members of the family transfer UDP, ADP, GDP or CMP linked sugars. The Glycos_transf_N region is flanked at the N-terminus by a signal peptide and at the C-terminus by Glycos_transf_1 (pfam00534). The eukaryotic glycogen synthases may be distant members of this bacterial family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCQ74891.1 | 2.89e-133 | 1 | 415 | 1 | 414 |
| CUW37378.1 | 7.52e-122 | 1 | 419 | 1 | 414 |
| BAE52896.1 | 3.44e-120 | 1 | 411 | 1 | 406 |
| AUG51867.1 | 4.61e-119 | 6 | 415 | 7 | 415 |
| QQP87941.1 | 1.73e-118 | 5 | 415 | 5 | 414 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2XCI_A | 3.96e-30 | 53 | 387 | 43 | 347 | Membrane-embeddedmonofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCI_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, substrate-free form [Aquifex aeolicus],2XCU_A Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_B Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_C Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus],2XCU_D Membrane-embedded monofunctional glycosyltransferase WaaA of Aquifex aeolicus, complex with CMP [Aquifex aeolicus] |
| 4BFC_A | 4.56e-23 | 228 | 391 | 39 | 203 | ChainA, 3-DEOXY-D-MANNO-OCTULOSONIC-ACID TRANSFERASE [Acinetobacter baumannii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q4UND5 | 6.61e-74 | 52 | 419 | 97 | 464 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) OX=315456 GN=waaA PE=3 SV=1 |
| Q92JE9 | 1.20e-70 | 51 | 419 | 96 | 464 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia conorii (strain ATCC VR-613 / Malish 7) OX=272944 GN=waaA PE=3 SV=1 |
| Q1RGU8 | 4.28e-69 | 4 | 415 | 3 | 413 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia bellii (strain RML369-C) OX=336407 GN=waaA PE=3 SV=1 |
| Q68XV7 | 2.64e-68 | 52 | 419 | 94 | 461 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia typhi (strain ATCC VR-144 / Wilmington) OX=257363 GN=waaA PE=3 SV=1 |
| Q9ZE58 | 1.08e-66 | 52 | 419 | 92 | 460 | 3-deoxy-D-manno-octulosonic acid transferase OS=Rickettsia prowazekii (strain Madrid E) OX=272947 GN=waaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
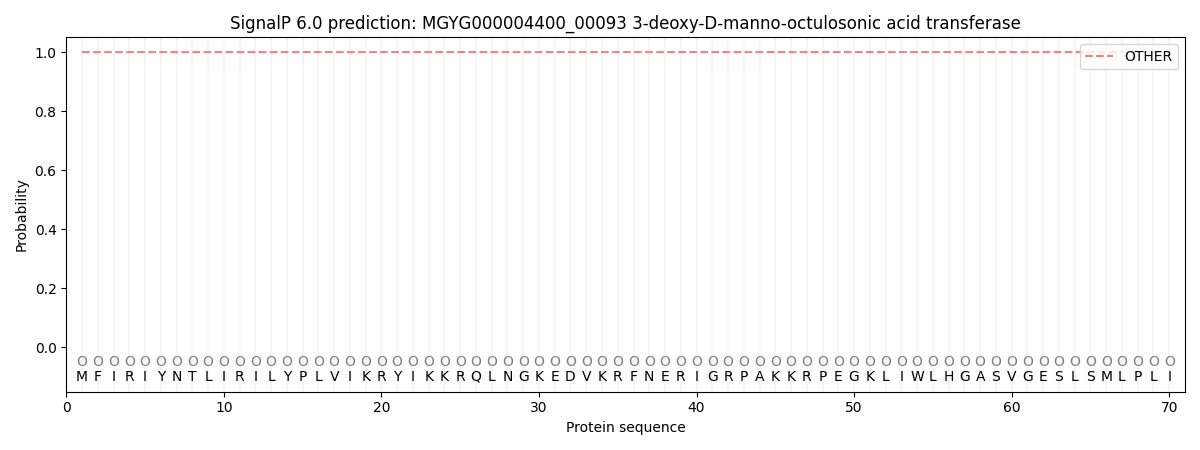
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999915 | 0.000091 | 0.000006 | 0.000000 | 0.000000 | 0.000000 |
